To learn more, please visit the YouTube Help Center: https://www.youtube.com/help
Get the latest international news and world events from around the world.
Is Quantum Entanglement Even Real?
To learn more, please visit the YouTube Help Center: https://www.youtube.com/help

New method turns ocean water into drinking water, without waste
The energy-efficient desalination system produces fresh water without chemical additives and transforms leftover salts into useful materials.

Meta wants to replace your mouse and keyboard with this bracelet
face_with_colon_three year 2025.
Researchers at Meta have developed a wristband that translates your hand gestures into commands to interact with a computer, including moving a cursor, and even transcribing your handwriting in the air into text. It could make today’s personal devices a lot more accessible to people with reduced mobility or muscle weakness, and even unlock new ways for people to control their gadgets effortlessly.

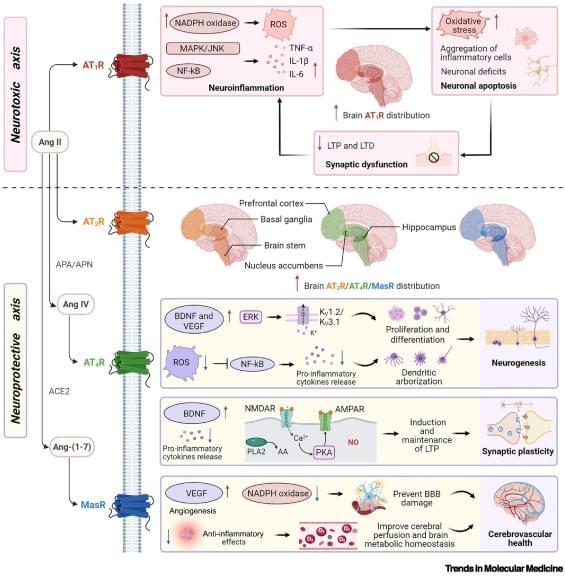
Renin–angiotensin system: a novel target for brain health
Emerging evidence highlights the brain renin–angiotensin system (RAS) as a key regulator of reward, memory, and stress. While these discoveries established the brain RAS as a promising therapeutic target for interventions in neurological and neuropsychiatric disorders, translational progress is hampered by the lack of an integrative mechanistic framework. Here, we consolidate accumulating evidence on the molecular and system-level roles of the brain RAS in reward, memory, and stress pathways, and its dual regulatory architecture. Pharmacological RAS modulation regulates domain-specific signaling in frontostriatal reward circuits, hippocampal–prefrontal memory networks, and frontolimbic fear networks. We evaluate the transdiagnostic therapeutic potential in neurological and neuropsychiatric disorders (e.g.

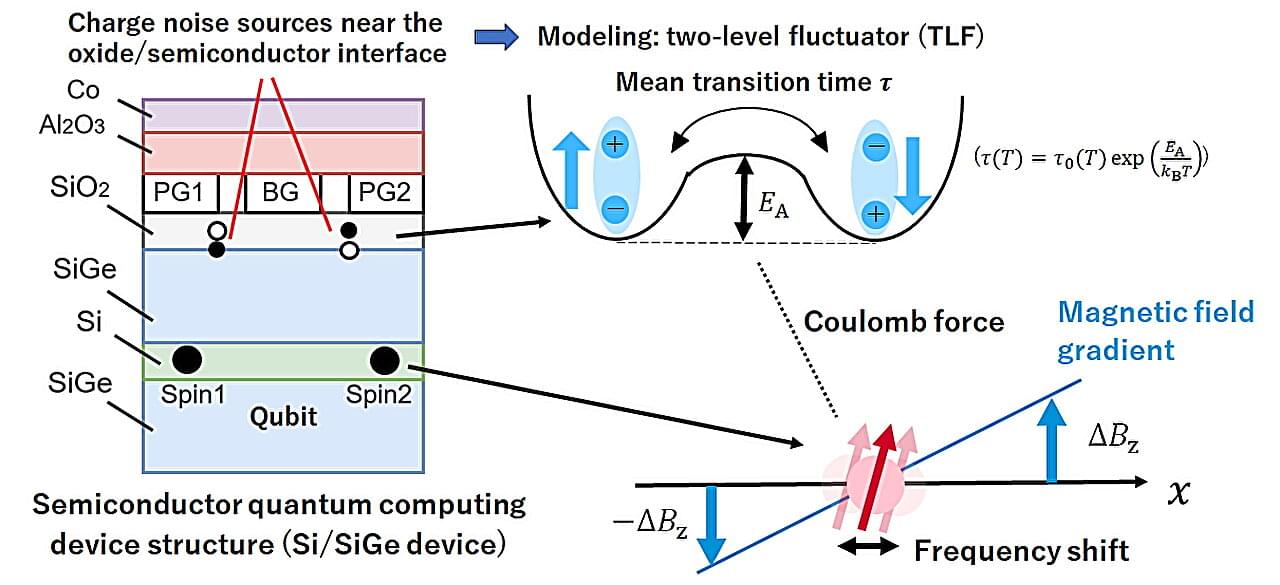
Scientists identify the origin of noise in spin qubit quantum processors
A spin qubit, in which quantum information is encoded in the spin state of an electron, is one of the most promising platforms for quantum computing. Spin qubits exhibit long coherence times and are compatible with advanced semiconductor manufacturing technologies. The leading implementation of spin qubits involves confined electrons inside quantum dots, a nanoscale semiconductor architecture that behaves like a controllable artificial atom. Recent advances have enabled high-fidelity operation of single- and two-qubit gates, exceeding the threshold required for certain surface code quantum error correction techniques.