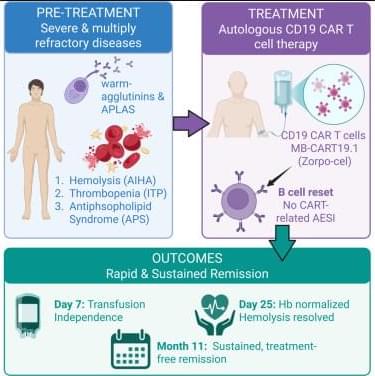
Cutting edge CAR-T cell therapy drives remission of three life-threatening autoimmune diseases in patient.
Med by Cell Press.
This single-patient case report expands the indications in which CD19 CAR-T cell therapy demonstrates unprecedented clinical efficacy, achieving sustained, treatment-free remission. Following B cell abrogation, AIHA is stopped, antiphospholipid antibodies are abrogated, and an underlying ITP is stabilized without any CAR-typical side effects.