Researchers relied on a newer gene-editing technique that may make it possible to engineer embryos, a prospect that has long alarmed bioethicists.

For 72 hours, enjoy 15% OFF on all Hoverpens with code PBS, or click the link https://noviumdesign.shop/PBS — Free shipping to most countries. Also on Amazon: https://noviumdesign.shop/crQH4A
We’ve been looking for messages from the stars ever since Frank Drake pointed the Green Bank radio telescope at Tau Ceti and Epsilon Eridany 65 years ago. He saw nothing that couldn’t be explained by natural causes. Nor have the much more extensive SETI surveys conducted since. So, maybe there are no alien signals to see. Or maybe we need to update how we search for them. We have, after all, learned an awful lot since 1960—both about the galaxy and about observing the galaxy.
Sign Up on Patreon to get access to the Space Time Discord! / pbsspacetime.
Check out the Space Time Merch Store https://www.pbsspacetime.com/shop.
Sign up for the mailing list to get episode notifications and hear special announcements! https://mailchi.mp/1a6eb8f2717d/space… the Entire Space Time Library Here: https://search.pbsspacetime.com/ Hosted by Matt O’Dowd Written by Matt O’Dowd Post Production by Leonardo Scholzer Directed by Andrew Kornhaber Associate Producer: Bahar Gholipour Executive Producer: Andrew Kornhaber Executive in Charge for PBS: Maribel Lopez Director of Programming for PBS: Gabrielle Ewing Assistant Director of Programming for PBS: Mike Martin Spacetime is a production of Kornhaber Brown for PBS Digital Studios. This program is produced by Kornhaber Brown, which is solely responsible for its content. © 2026 PBS. All rights reserved. End Credits Music by J.R.S. Schattenberg: / multidroideka Space Time Was Made Possible In Part By: Big Bang Alexander Tamas Filip Rolenec Juan Benet Kenneth See Mark Rosenthal Matthew Ocko Morgan Hough Peter Barrett Vinnie Falco Daniel Muzquiz Quasar Ethan Cohen Glenn Sugden Grace Biaelcki Justin Lloyd Mark Heising Rad Antonov Shaun Williams Stephen Wilcox Tristan Lucian Claudius Aurelius Tyacke Hypernova Alex Kern Ben Delo Chuck Zegar Dean Galvin Donal Botkin Gregory Forfa Jeff White John R. Slavik Massimiliano Pala PAUL C PEDERSEN Scott Gorlick Scott Gray Spencer Jones Vlad Shipulin Zachary Haberman Антон Кочков Gamma Ray Burst Alex Gan aaron pinto Almog Cohen Anthony Leon Arko Provo Mukherjee Ayden Miller Bradley Jenkins Bradley Ulis Brandon Lattin Brian Cook Chris Liao Christopher Wade Chuck Lukaszewski Collin Dutrow Craig Falls Craig Stonaha Dan Warren Daniel Donahue Daniel Jennings Darrell Stewart David Giltinan David Johnston Doyle Vann Eric Kiebler Eric Raschke Eric Schrenker Faraz Khan Frederic Simon gmmiddleton Harsh Khandhadia Isaac Suttell James Trimmier Jason Bowen Jeb Campbell Jeff Harris Jeremy Soller Jerry Thomas jim bartosh John Anderson John De Witt John Funai John H. Austin, Jr. Joseph Salomone Junaid Ali Kacper Cieśla Kane Holbrook Kent Durham Koen Wilde Kyle Atkinson Lori Ferris Marcelo Garcia Marion Lang Mark Daniel Cohen Mark Delagasse Matt Kaprocki Matt Quinn Matthew Johnson Michael Barton Michael Clark Michael Lev Michael Purcell Mikk Mihkel Nurges Nick Hoffenstoffer III Nicolas Katsantonis Onemind Param Saxena Paul Wood Rad Antonov Reuben Brewer Richard Steenbergen Robert DeChellis Ross Kennedy Ross Story Russell Moore SamSword Sandhya Devi Sean Owen Shane Calimlim SilentGnome Terje Vold Thomas Dougherty Todd J Lerner Tybie Fitzhugh Zac Sweers.
Search the Entire Space Time Library Here: https://search.pbsspacetime.com/
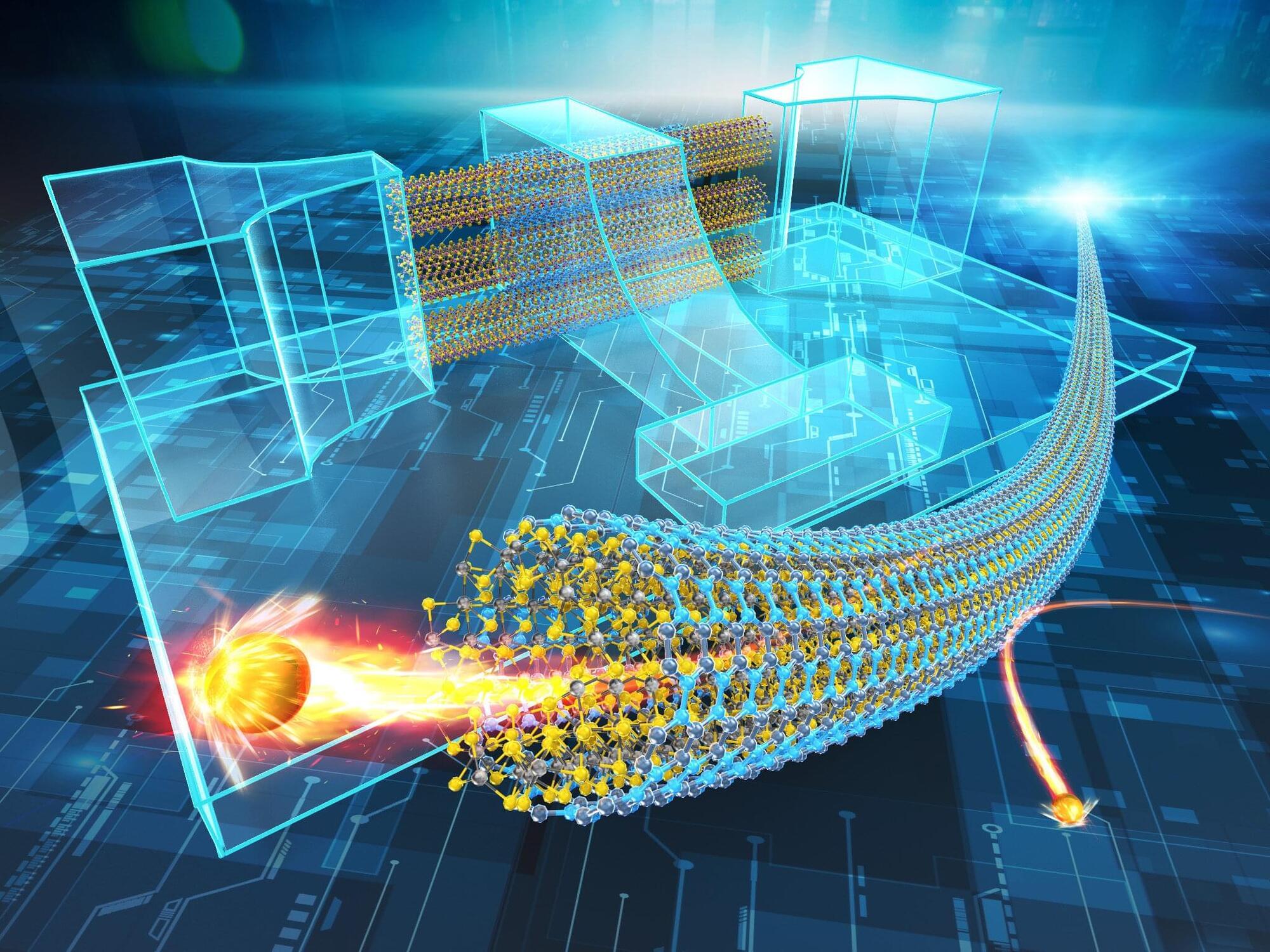
Researchers in Japan have created some of the world’s smallest semiconducting nanotubes, structures 100,000 times thinner than a human hair. By growing molybdenum disulfide inside protective tubes of boron nitride, the researchers, including those from the University of Tokyo, produced highly uniform tubes just 1 nanometer wide, a scale at which it’s difficult to make stable nanotube structures. The work confirms decades-old theoretical predictions about how these ultrafine materials behave and could also provide a new route toward miniaturized electronic devices.
The research is published in the journal Science.
A few years ago, carbon nanotubes were attracting a lot of press attention. But there’s a new contender in the ring, and it offers some advantages over its carbon counterpart that could tempt engineers to design products around it.
In this video, we explore the incredible and terrifying world of nano-weapons — microscopic machines designed for the battlefields of the future. From invisible drones to molecular-level assassins, nanotechnology is revolutionizing modern warfare in ways the world has never seen before. Discover how these tiny machines can spy, sabotage, and even kill at the atomic scale. We’ll uncover real-world research, secret military projects, and the ethical dangers behind the next generation of warfare. The rise of nano-weapons could change the balance of global power forever — but are we ready for what’s coming? Watch till the end to understand the full potential and risks of these microscopic war machines.
Hashtags (12):
#NanoWeapons #FutureWarfare #Nanotechnology #MilitaryTech #WarInnovation #ScienceAndTechnology #NanoMachines #InvisibleWar #BattlefieldOfTomorrow #DefenseTechnology #AIWarfare #FutureScience.
Keywords (23):
nano weapons, microscopic machines, future warfare, military technology, nanotechnology, nano drones, nano robots, molecular weapons, defense innovation, secret military research, nano science, nano warfare, nano army, advanced technology, invisible battlefield, modern weapons, AI warfare, nanobots, nano defense, future military, scientific discovery, advanced warfare, nano.
NASA’s Nancy Grace Roman Space Telescope is gearing up to unlock the deepest secrets of the universe with three groundbreaking surveys. Designed with input from over a thousand scientists worldwide, Roman’s missions will map billions of galaxies, capture the dynamic dance of cosmic phenomena like
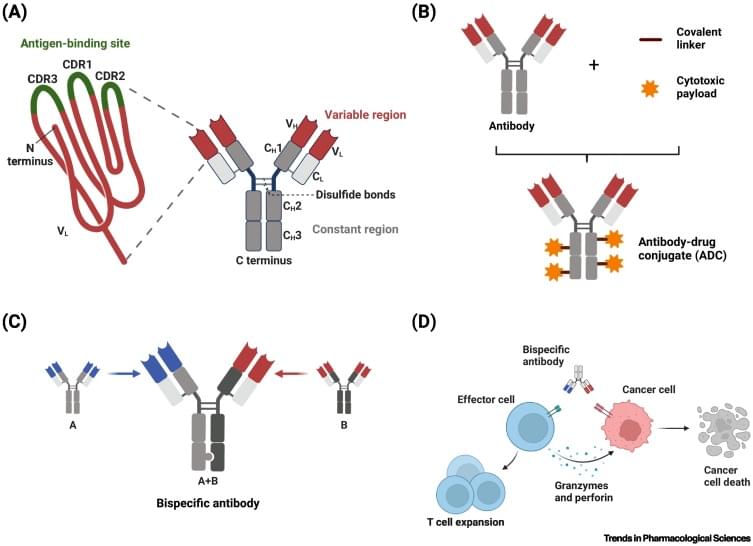
Antibodies in oncology are being equipped with toxic cargoes and effector functions that can kill cells at very low concentrations. A key challenge is that most targets on cancer cells are also present on at least some healthy cells. Shared targets can result in off-tumor binding and compromise the safety and potential of therapeutic candidates. In this review, we survey strategies that can help direct biologics to cancer sites more selectively. These strategies are becoming increasingly feasible thanks to advances in molecular design and engineering. The objective is to create therapeutics that exploit changes in cancer and leverage the human body infrastructure, enabling therapeutics that discriminate not just self from non-self but diseased from healthy tissue.
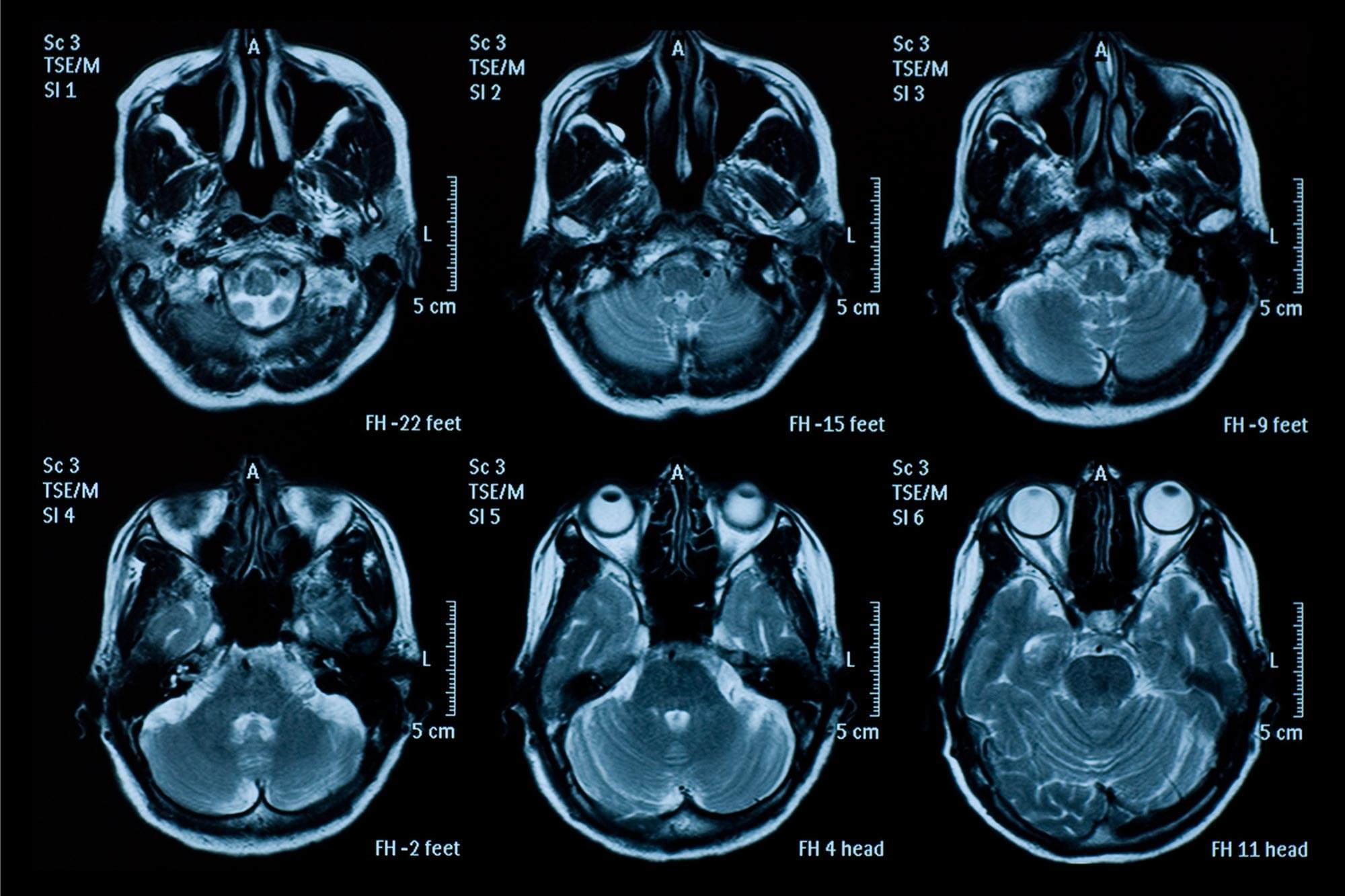
Even mild concussion can cause long-lasting effects to the brain, according to researchers at the University of Cambridge. Using data from a Europe-wide study, the team has shown that for almost a half of all people who receive a knock to the head, there are changes in how regions of the brain commu
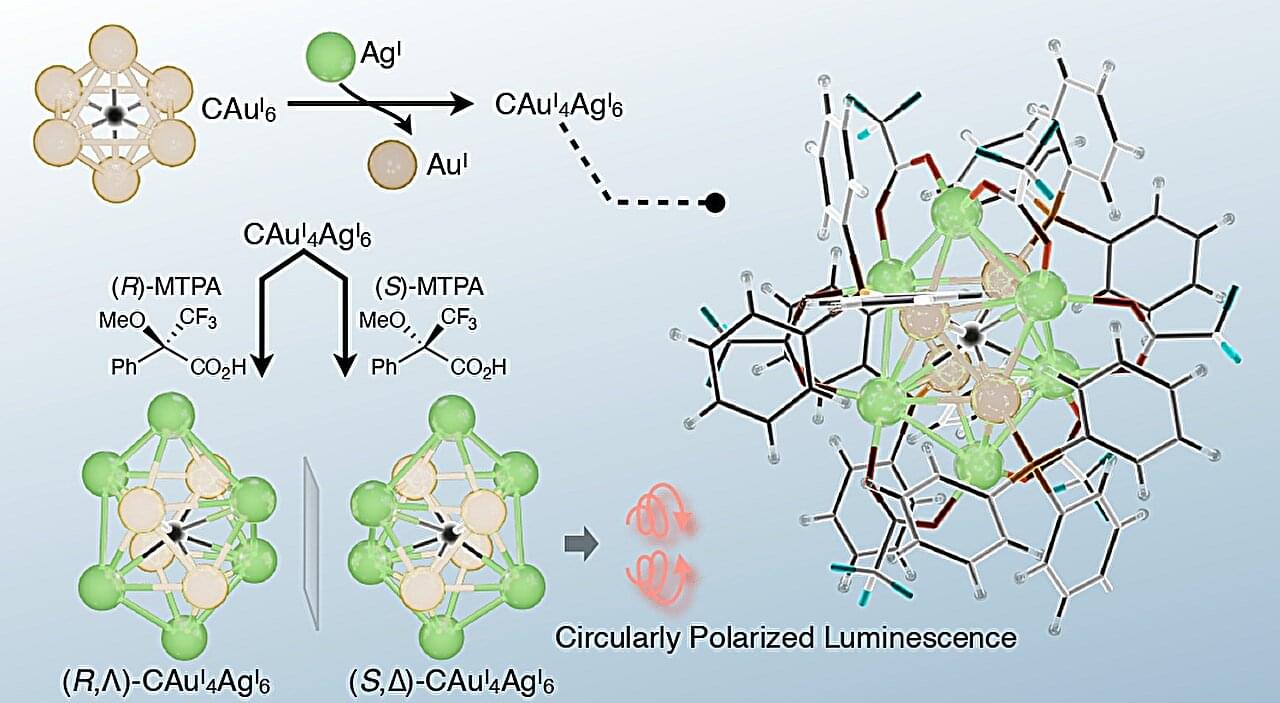
Metal cluster molecules are discrete compounds containing multiple metal atoms held together by metal–metal and metal–ligand bonding. They serve as excellent candidates for catalysts, biosensors, and even for drug development. Developing atomic-level molecular editing methods for such metal clusters remains an important challenge and represents a promising strategy for expanding their structural and functional diversity. Such approaches can enable structure-specific properties, high near-infrared (NIR) photoluminescence quantum yields, and unique reactivities and electronic structures.
Alloying is a powerful method for achieving this goal. In this regard, a key challenge is asymmetric alloying, which introduces asymmetry into the metal cluster by selectively placing heterometal atoms at nonequivalent sites, desymmetrizing the cluster and therefore imparting chirality-associated functionality.
Moreover, highly selective asymmetric synthesis methods for heterometallic clusters are expected to contribute significantly to the development of chiroptical materials. However, methods capable of achieving such controlled asymmetric synthesis have rarely been reported.


Storm Dave, which swept across northern Europe over the Easter weekend, is an example of what new research from the University of Gothenburg has revealed. Spring storms forming over the North Atlantic have become more common than they were 80 years ago, and this is due to climate change.
In the Northern Hemisphere, storm seasons follow a seasonal cycle. Storms are weakest and least frequent in summer and most intense in winter. As a result of global warming, storm patterns and their course have changed, and several studies have indicated that winter storms appear to be occurring more frequently and with even greater intensity.