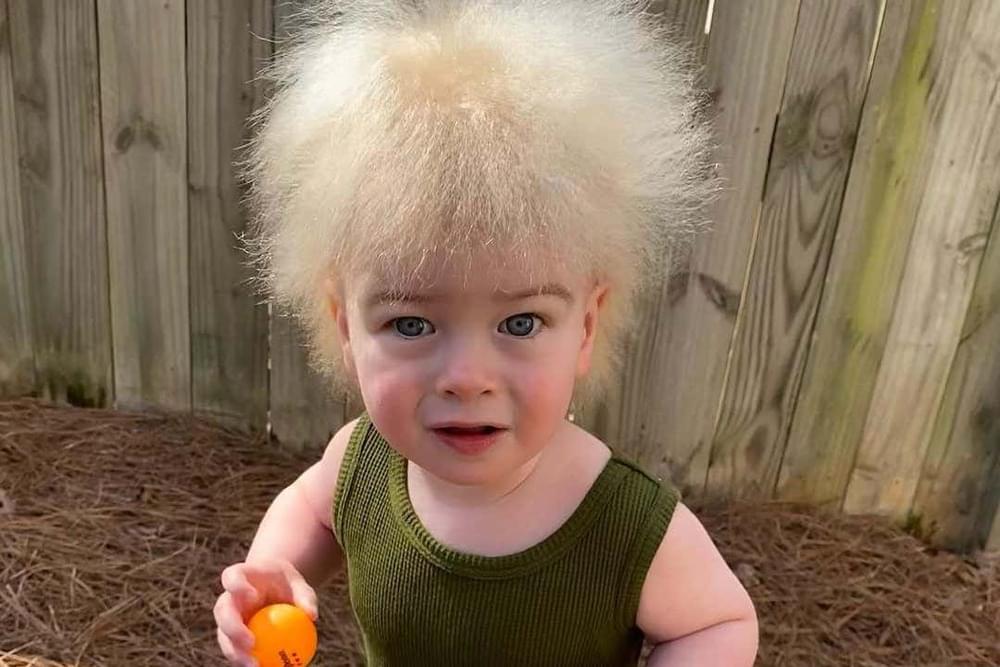
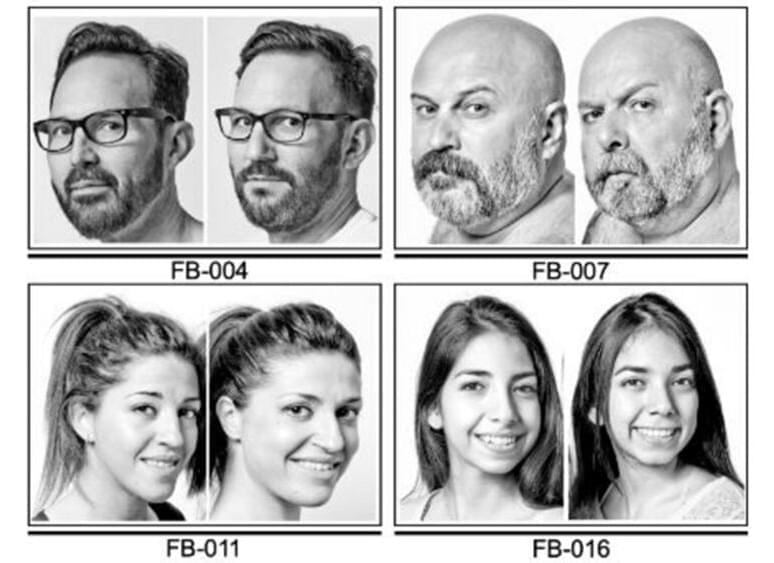
Most common genetic causes of a rare condition called “uncombable hair syndrome” have been found.



Foresight Existential Hope Group.
Program & apply to join: https://foresight.org/existential-hope/
In the Existential Hope-podcast (https://www.existentialhope.com), we invite scientists to speak about long-termism. Each month, we drop a podcast episode where we interview a visionary scientist to discuss the science and technology that can accelerate humanity towards desirable outcomes.
Xhope Special with Foresight Fellow Morgan Levine.
Morgan Levine is a ladder-rank Assistant Professor in the Department of Pathology at the Yale School of Medicine and a member of both the Yale Combined Program in Computational Biology and Bioinformatics, and the Yale Center for Research on Aging. Her work relies on an interdisciplinary approach, integrating theories and methods from statistical genetics, computational biology, and mathematical demography to develop biomarkers of aging for humans and animal models using high-dimensional omics data. As PI or co-Investigator on multiple NIH-, Foundation-, and University-funded projects, she has extensive experience using systems-level and machine learning approaches to track epigenetic, transcriptomic, and proteomic changes with aging and incorporate.
this information to develop measures of risk stratification for major chronic diseases, such as cancer and Alzheimer’s disease. Her work also involves development of systems-level outcome measures of aging, aimed at facilitating evaluation for geroprotective interventions.
Existential Hope.
A group of aligned minds who cooperate to build beautiful futures from a high-stakes time in human civilization by catalyzing knowledge around potential paths to get there and how to plug in.
Follow us!
Twitter: https://twitter.com/HopeExistential.
Instagram: https://www.instagram.com/existential.hope/
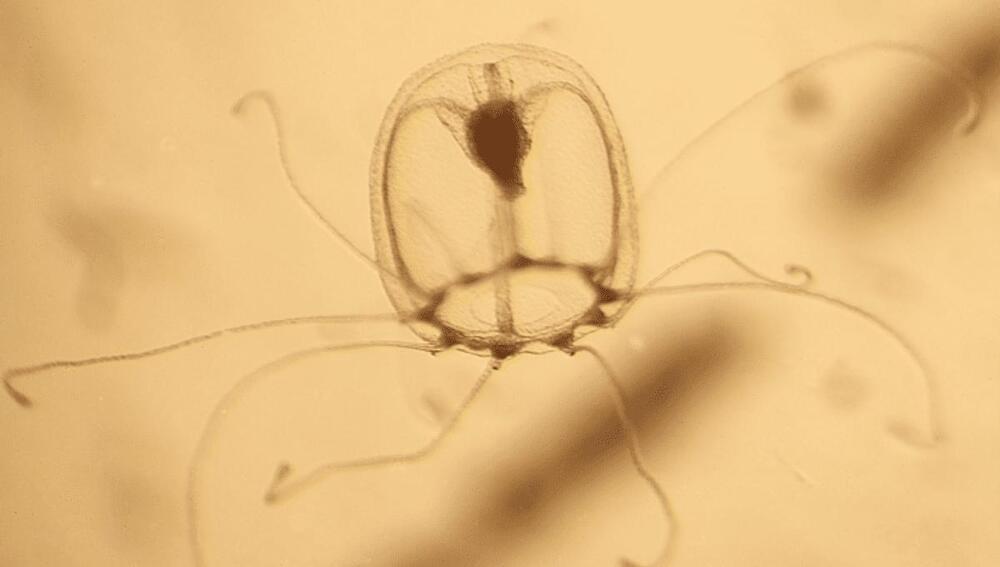
Immortality exists – but to get it, you need to be a jellyfish, not a god or a vampire. Moreover, only one species of cnidarian, Turritopsis dohrnii, is known to have found the secret of eternal life. Geneticists hope comparing T. dornii’s DNA with its close relative, T. rubra, will help us understand the aging process and how to evade it.
Turritopsis are warm water jellyfish half a centimeter (0.2 inches) long. At least three species of hydra have the capacity to age backwards like Benjamin Button, going from adult to juvenile stage, before eventually growing up again. However, two of these can only go from the hydra equivalent of adolescent to child; like the victim in some uncensored fairytale, sexual reproduction locks them into adulthood. T. dohrnii, on the other hand, appears able to go from its free-floating adult stage to bottom-living polyp, known as life cycle reversal (LCR), as many times as it wants.
A paper in the journal Proceedings of the National Academy of Sciences provides a comparison of T. dorhnii and T. rubra in the hope the differences will prove enlightening, throwing in a few more distantly related types of cnidarians as well.
Managing Complex Transplant (and Xenotransplant) Cases Globally — Dr Robert A Montgomery, MD, DPhil, FACS, Director, NYU Langone Health
Dr. Robert A. Montgomery, MD, DPhil, FACS, (https://nyulangone.org/doctors/1467404137/robert-montgomery) is the Director of the NYU Langone Transplant Institute, and Chair and a Professor in their Department of Surgery, where he oversees a diverse team of medical and surgical specialists who provide a wide variety of surgery and transplantation services including bone marrow, heart, kidney, liver, lung, and facial transplantation.
Dr. Montgomery received his Doctor of Medicine with Honor from the University of Rochester School of Medicine, his Doctor of Philosophy from Balliol College, The University of Oxford, England in Molecular Immunology, and completed his general surgical training, multi-organ transplantation fellowship, and postdoctoral fellowship in Human Molecular Genetics at Johns Hopkins.
For over a decade Dr. Montgomery served as the Chief of Transplant Surgery and the Director of the Comprehensive Transplant Center at Johns Hopkins.
Dr. Montgomery was part of the team that developed the laparoscopic procedure for live kidney donation, a procedure that has become the standard throughout the world. He and the Hopkins team conceived the idea of the Domino Paired Donation (kidney swaps), the Hopkins protocol for desensitization of incompatible kidney transplant patients, and performed the first chain of transplants started by an altruistic donor. He led the team that performed the first 2-way, 3-way, 4-way, 5-way, 6-way, and 8-way domino paired donations, and in the first 10-way open chain donation.

Join us on Patreon!
https://www.patreon.com/MichaelLustgartenPhD
TruDiagnostic Discount Link (Epigenetic Testing)
CONQUERAGING!
https://bit.ly/3Rken0n.
Bristle Discount Link (Oral microbiome quantification):
ConquerAging15
https://www.bmq30trk.com/4FL3LK/GTSC3/
Cronometer Discount Link (Daily diet tracking):
https://shareasale.com/r.cfm?b=1390137&u=3266601&m=61121&urllink=&afftrack=
Support the channel with Buy Me A Coffee!
https://www.buymeacoffee.com/mlhnrca

The groundbreaking research that established the connection between Alzheimer’s.
Alzheimer’s disease is a disease that attacks the brain, causing a decline in mental ability that worsens over time. It is the most common form of dementia and accounts for 60 to 80 percent of dementia cases. There is no current cure for Alzheimer’s disease, but there are medications that can help ease the symptoms.
The more we understand how cells produce shape and form, the more inadequate the idea of a genomic blueprint looks by Philip Ball + BIO.
