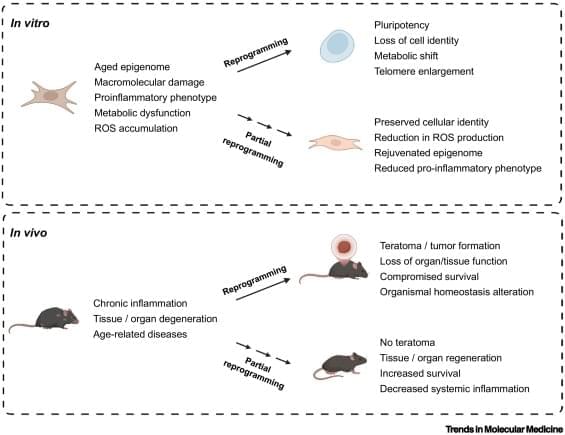
Aging, once viewed as an irreversible process, is now considered a modifiable process. Recent advances in cellular reprogramming reveal that transient expression of reprogramming factors can reverse molecular hallmarks of aging while preserving somatic cell identity. This ‘partial reprogramming’ rejuvenates tissues, restores regenerative capacity, and, in some models, extends lifespan without the tumorigenic risks of full dedifferentiation. In this review, we summarize genetic and chemical strategies for partial reprogramming, discuss their tissue-specific effects in vivo, and evaluate their implications for tissue regeneration and age-related disease. We further examine key challenges for clinical translation, including safety, delivery strategies, and temporal control of reprogramming.