This website uses a security service to protect against malicious bots. This page is displayed while the website verifies you are not a bot.


A personalized vaccine to treat glioblastoma, a fast-growing and incurable brain cancer that affects four in 100,000 people in the U.S., is safe and elicits robust and broad immune responses that appears to increase recurrence-free survival in a subset of patients after surgery, according to an early-stage clinical trial co-led by researchers at Washington University School of Medicine in St. Louis.
In patients with an especially aggressive form of glioblastoma, the vaccine caused no serious side effects and prolonged patients’ overall survival compared to historical outcomes after standard-of-care surgery and chemo-radiotherapy. One long-term survivor remains recurrence-free nearly five years later.
The results of the phase 1 trial, conducted at Siteman Cancer Center, based at Barnes-Jewish Hospital and WashU Medicine, were published May 12 in Nature Cancer. The study was led jointly by Mass General Brigham and Geneos Therapeutics, a Philadelphia-based biotechnology company.
“We are extremely encouraged by these results,” said Tanner M. Johanns, MD, PhD, lead author of the study and an assistant professor in the Division of Oncology in the John T. Milliken Department of Medicine at WashU Medicine. “This kind of vaccine is a first for glioblastoma, and it is exciting to think how we can leverage this individualized therapeutic DNA cancer vaccine platform to make a positive impact on the lives of patients who are fighting this disease. Additionally, combination therapies leveraging this personalized platform are currently being investigated at WashU to test if outcomes may be improved further.”
Abstract: Nature Cancer
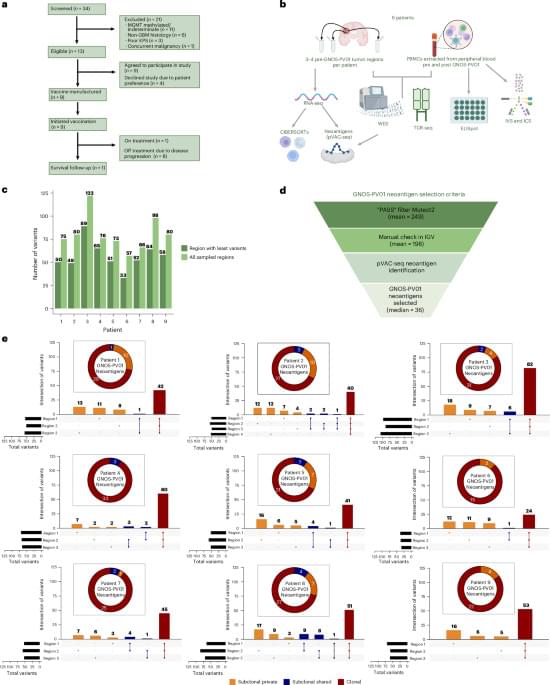
Johanns and colleagues report the results (including safety, efficacy and immunogenicity) of a phase 1 clinical trial of a DNA-based personalized therapeutic cancer vaccine administered following surgical resection and radiation in patients with MGMT unmethylated glioblastoma.
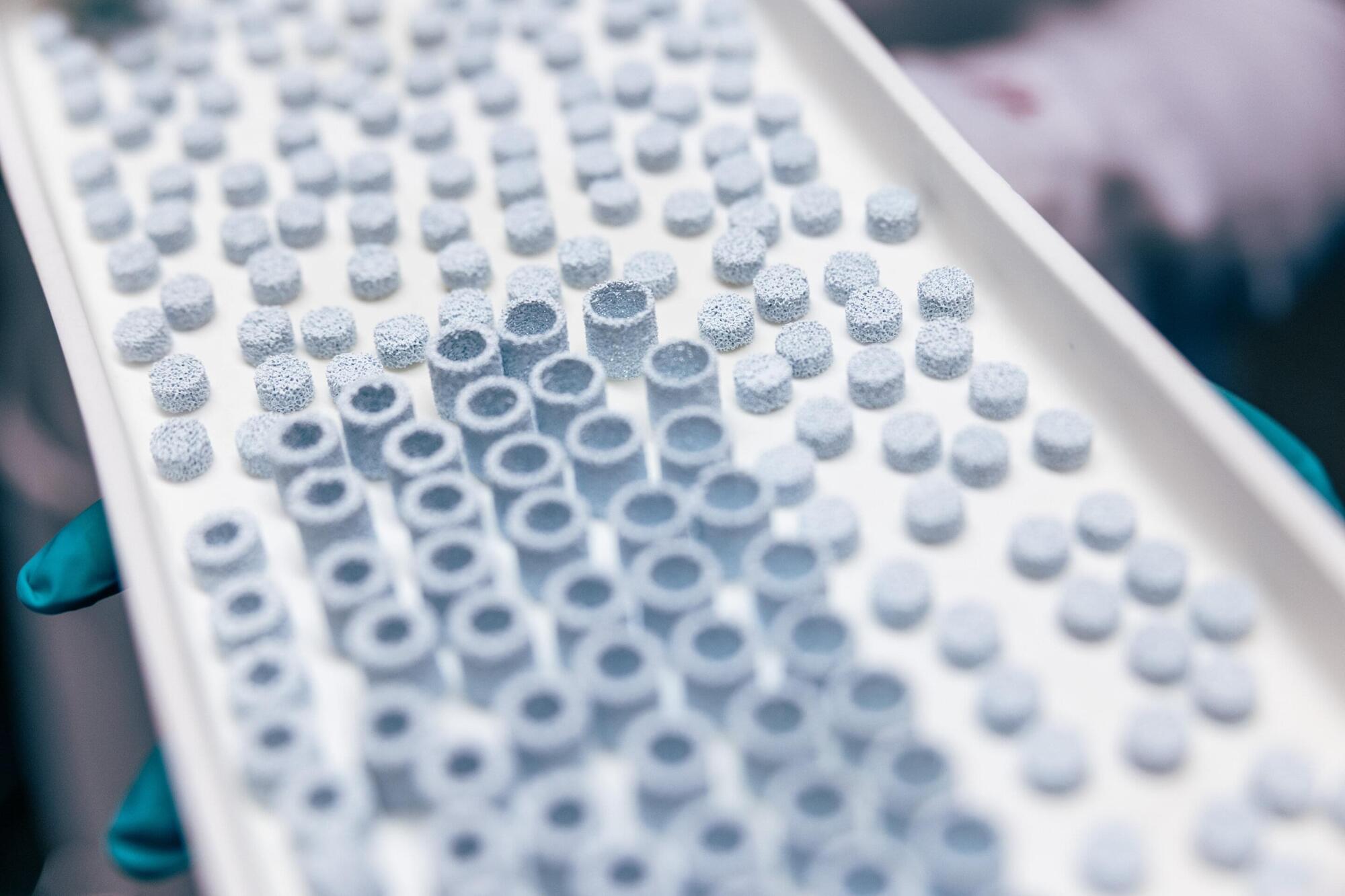

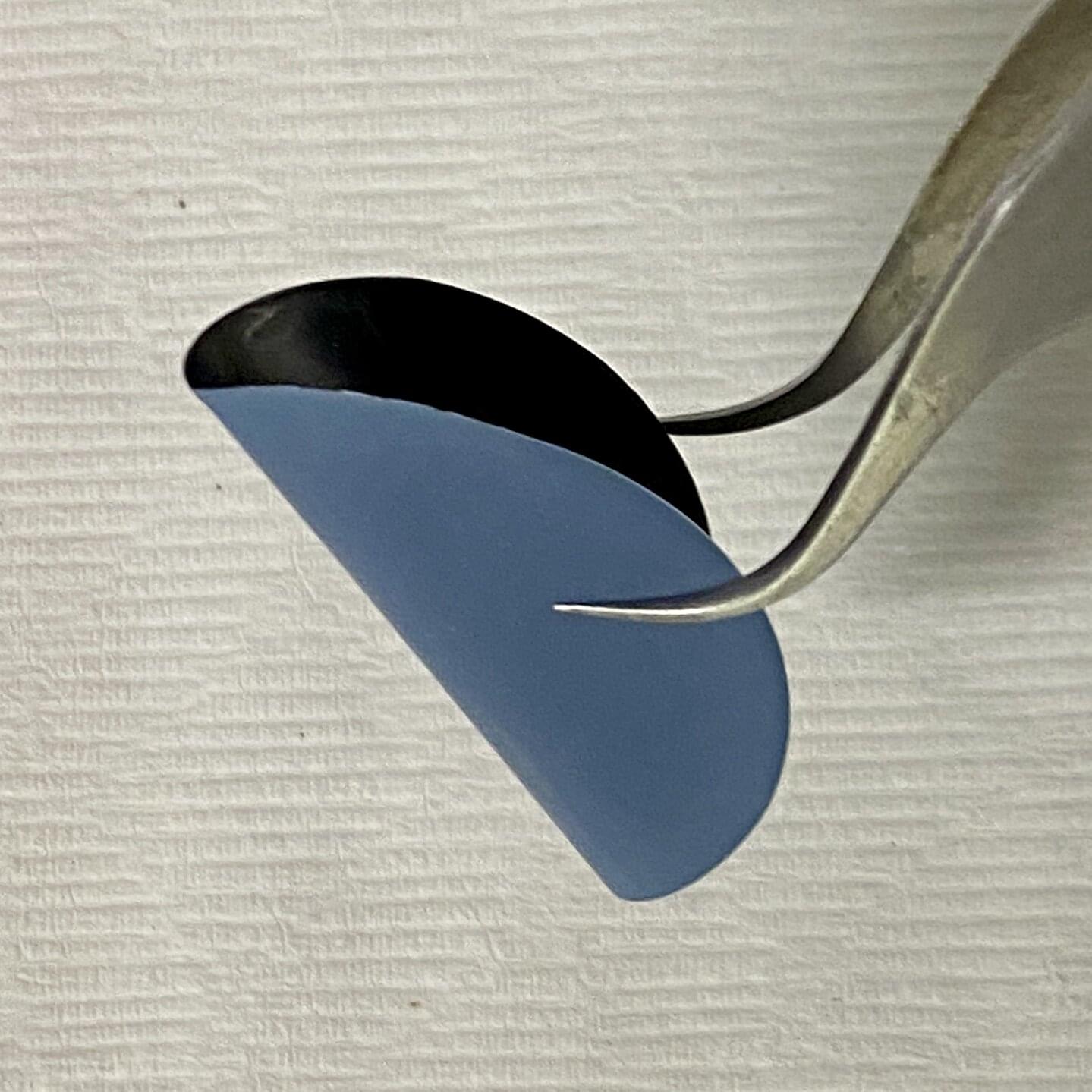
Researchers at Tampere University, Finland, have developed a groundbreaking 3D-printed ceramic implant material that closely mimics real human bone. The findings advance the development of personalized bone regeneration and may lead to more effective and accessible treatments for bone defects.
The research article, titled “Biomimetic bone calcium phosphate-based scaffolds fabricated via ceramic vat photopolymerization: Effect of porosity, sintering temperature, mineralogical phases and trace elements on the osteogenic potential,” was published in Materials Today Bio.
Bone grafting is the second most common tissue transplantation procedure worldwide, with more than 2 million operations performed annually. Current treatments often rely on bone taken from the patient or a donor, approaches that are limited in availability and may involve additional surgery, lengthy recovery times and complications. As populations age, the need for safer and more effective alternatives is growing rapidly.
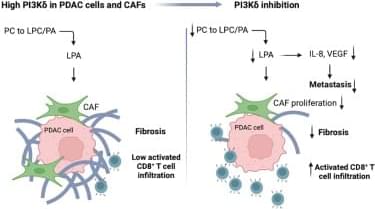
In this study, we identify an unexpected and previously unrecognized role for PI3Kδ in promoting stromal fibrosis in PDAC, expanding its known function beyond immune regulation. Through mechanistic and preclinical studies, we show that PI3Kδ controls the biosynthesis of LPA in cancer cells and stromal fibroblasts, establishing an immunometabolic axis that sustains both fibrosis and immune evasion in PDAC.
Strikingly, PI3Kδ inhibition alone was sufficient to suppress tumor growth, reduce fibrosis, restore antitumor immune responses, and prolong survival across multiple PDAC models. Dual inhibition of PI3Kδ and ATX produced additive effects on stromal remodeling and immune activation, significantly enhancing responsiveness to chemotherapy and PD-1 blockade. These findings position PI3Kδ as a central regulator of the PDAC tumor microenvironment and highlight its therapeutic targeting, alone or in combination, as a promising strategy to treat PDAC.
A pioneering treatment which could slow or reverse liver failure and offer a potential alternative to liver transplants has shown positive results in a medical trial.
70% of end-stage liver disease patients who were treated with macrophage cell therapy in the MATCH trial did not need a liver transplant after four years, compared with just 40% who didn’t receive the treatment.
The cell therapy takes immune cells from the patients’ blood and turns them into mature macrophages – a white blood cell – which is then re-injected back into the patient. The macrophages travel to the liver, where they break down scar tissue, reduce inflammation, and encourage the growth of healthy liver cells.
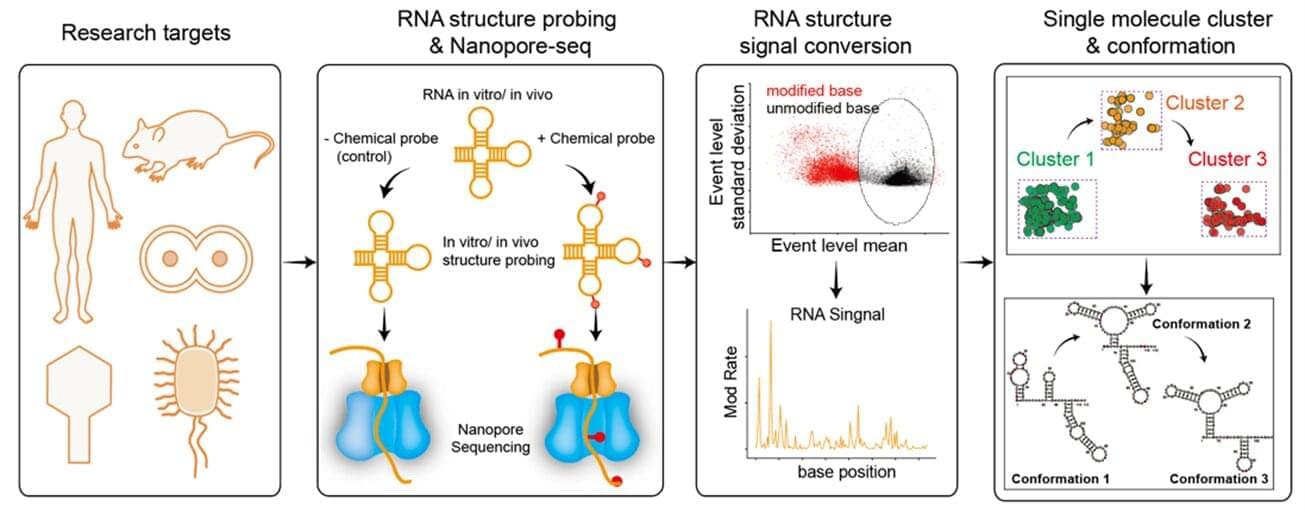
Researchers from A*STAR Genome Institute of Singapore (A*STAR GIS) have developed a new method to study individual RNA molecules and reveal how their structures influence gene regulation, a fundamental process that affects how cells function in health and disease. Their work was published in Nature Methods.
RNA is best known for carrying genetic instructions from DNA to make proteins. However, RNA does more than act as a messenger. Like a string that can bend, fold and interact with other molecules, RNA can adopt different shapes that affect how it behaves in the cell. These shapes can influence how efficiently proteins are produced, how long RNA molecules last, and how diseases such as viral infections progress.
Until now, studying these structures in detail has been difficult because RNA is highly flexible and dynamic. Most existing methods only provide an average picture across many RNA molecules, making it harder to see how individual RNA molecules may fold differently, even when they come from the same gene.

Engineers have developed a new ultra-thin membrane that allows fuel cells to operate more efficiently at high temperatures by enabling proton transport without water, overcoming a key limitation in clean energy technologies.
The breakthrough, reported in Science Advances, could expand the use of fuel cells in transport, heavy industry, and future clean energy systems.
Fuel cells convert chemical energy directly into electricity, producing water and heat as the main by-products. They are already used in hydrogen-powered vehicles, backup power systems for hospitals and data centers, and space missions where lightweight, reliable energy is essential.

A routine experiment with a new single-cell DNA sequencing method turned into a surprising scientific twist when researchers stumbled upon a bizarre genetic code in a microscopic pond organism. Instead of following the near-universal “rules” of life, this newly identified protist rewrites how genes signal their end. This unexpected discovery challenges long-held assumptions about how genetic translation works and hints that nature may be far more flexible—and mysterious—than scientists realized.