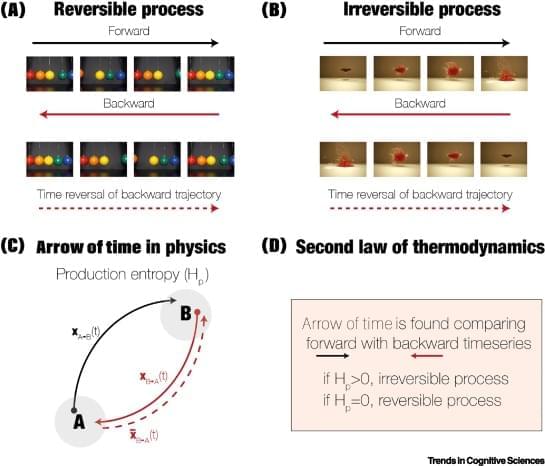
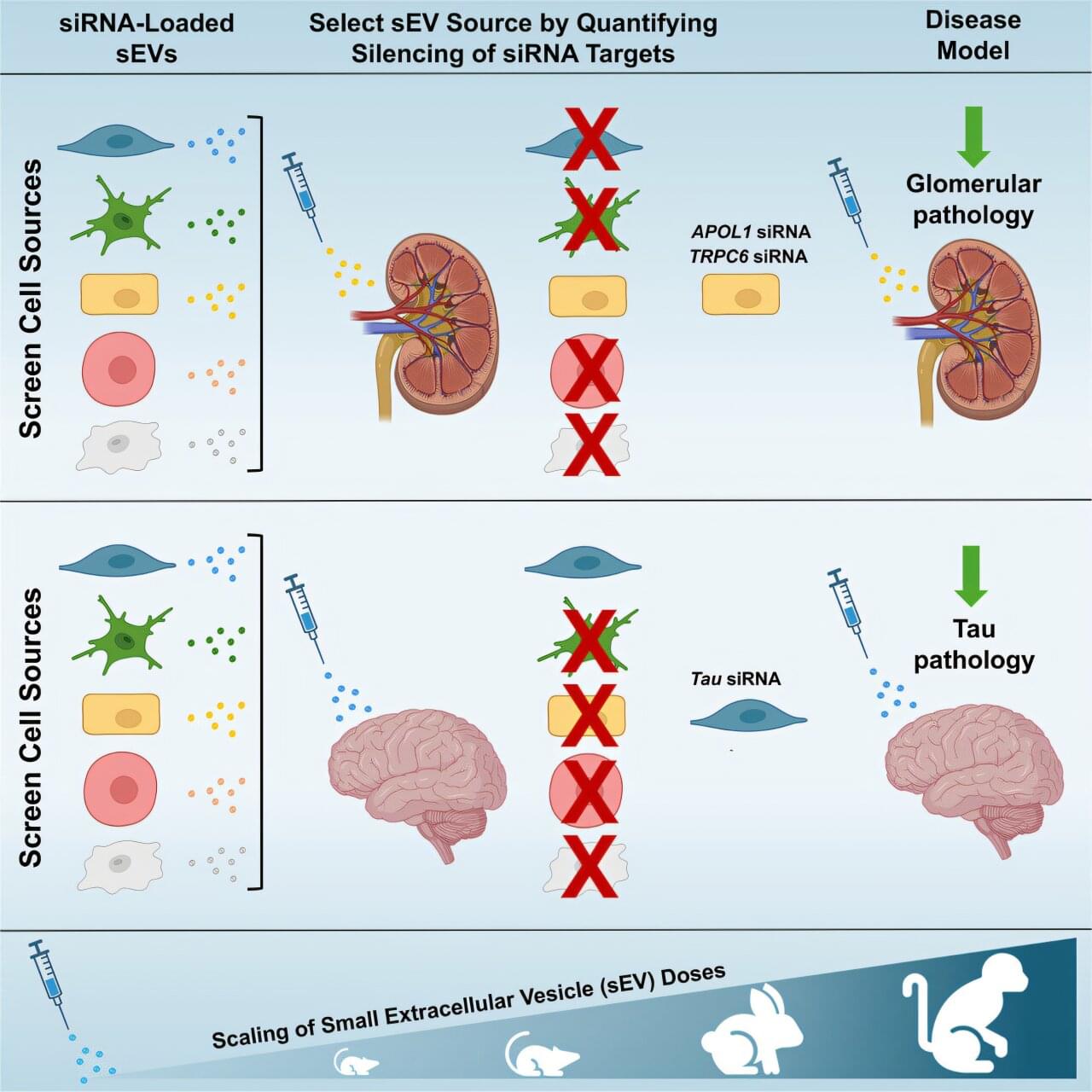
To not only survive, but also thrive, the brain must efficiently orchestrate distributed computation across space and time. This requires hierarchical organisation facilitating fast information transfer and processing at the lowest possible metabolic cost. Quantifying brain hierarchy is difficult but can be estimated from the asymmetry of information flow. Thermodynamics has successfully characterised hierarchy in many other complex systems. Here, we propose the ‘Thermodynamics of Mind’ framework as a natural way to quantify hierarchical brain orchestration and its underlying mechanisms. This has already provided novel insights into the orchestration of hierarchy in brain states including movie watching, where the hierarchy of the brain is flatter than during rest. Overall, this framework holds great promise for revealing the orchestration of cognition.