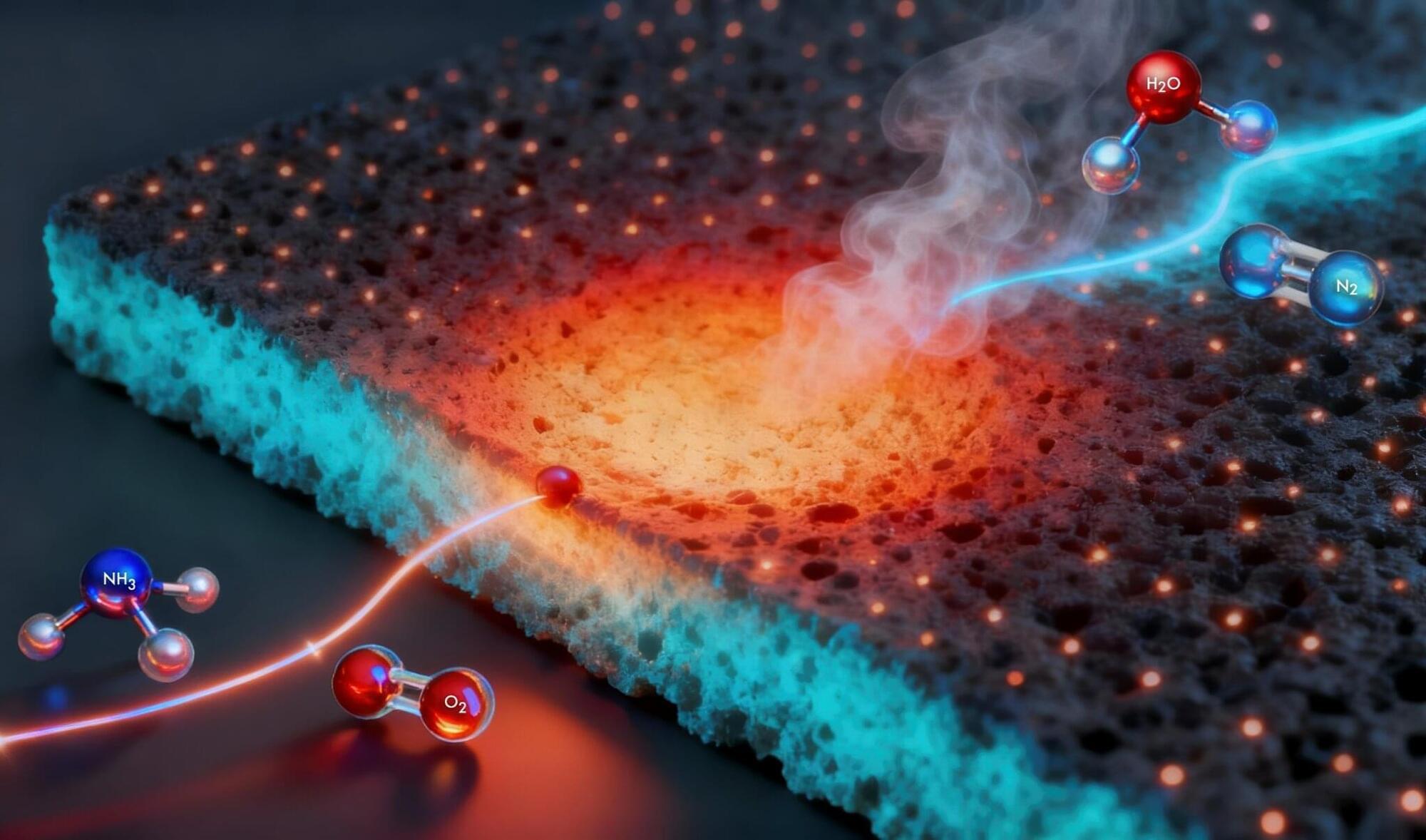
A single-atom platinum catalyst lights ammonia at 200 °C and keeps it burning steadily at 1,100 °C with low NOx, generating high-grade, carbon-free heat for steel, cement and chemicals.

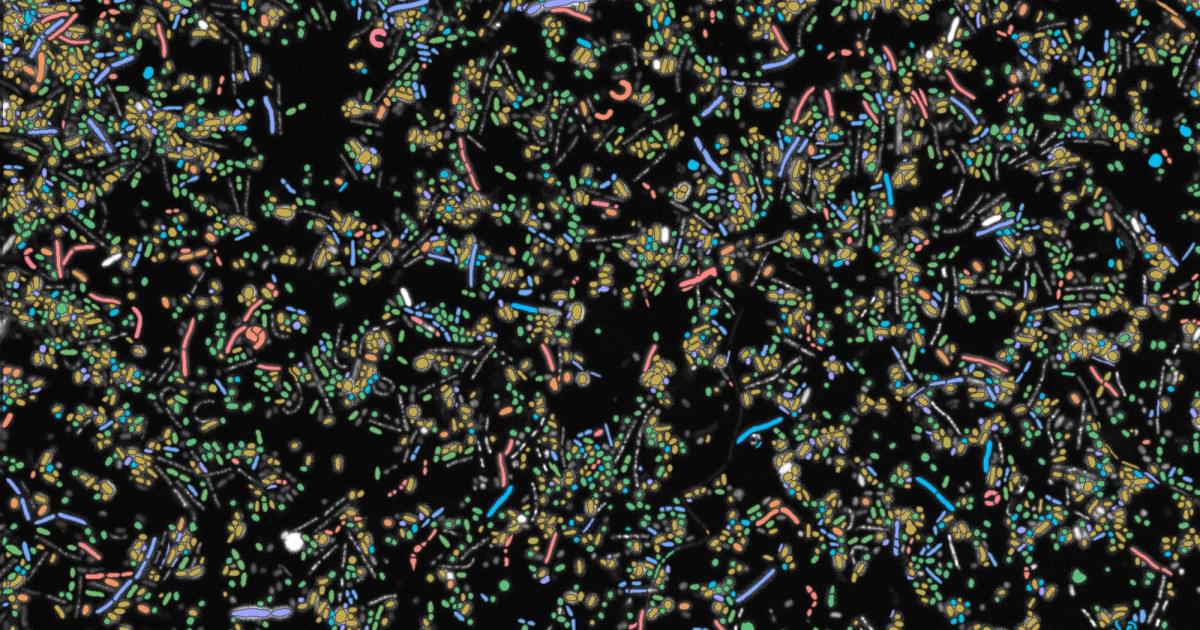
Kanvas looks amazing! They’re systematically deciphering microbiomes and developing clinical-stage interventions to improve patient outcomes in oncology and beyond. Very impressive! I’m also especially interested in their approach to maternal environmental enteric dysfunction (EED), which apparently affects 150M people!
Ever since the genomics revolution revealed how reliant the human organism is on its microscopic microbial cohabitants, the microbiome has been medicine’s most elusive frontier, promising better health if only we could untangle the trillions of interactions that influence nearly every facet of our physiology. But until now, effective medicines that harness the microbiome have been rare. Because of the diversity of microbial species and the complexity of host-to-microbe interactions, as well as the lack of a reliable, easily manufactured drug modality (the package that delivers a medicine’s therapeutic effect), the microbiome has been hard to treat, despite its importance to functions like immune response. Microbiome science has disappointed patients, doctors, founders, and investors.
That’s why DCVC is so excited about the cascade of recent developments at Kanvas Biosciences, which is moving the field beyond descriptive profiling of the microbiome to translating comprehensive biochemical insights into clinically useful products. In the past few weeks, the Princeton-based spatial biology company has kicked off a Phase 1 clinical trial for its first drug candidate, secured significant new backing from the Gates Foundation (closing a $48 million Series A financing, bringing Kanvas’s total funding to $78 million), and bolstered its scientific leadership by adding one of the most respected names in bioengineering to its board.
Clinical milestone
The most significant milestone in Kanvas’s evolution is the dosing of the first patients in a Phase I clinical trial for KAN-4. This live biotherapeutic product (LBP), resembling an ordinary pill, treats the colitis that many cancer patients develop after receiving immune checkpoint inhibitors (ICIs), allowing them to remain on the life-saving therapy longer.
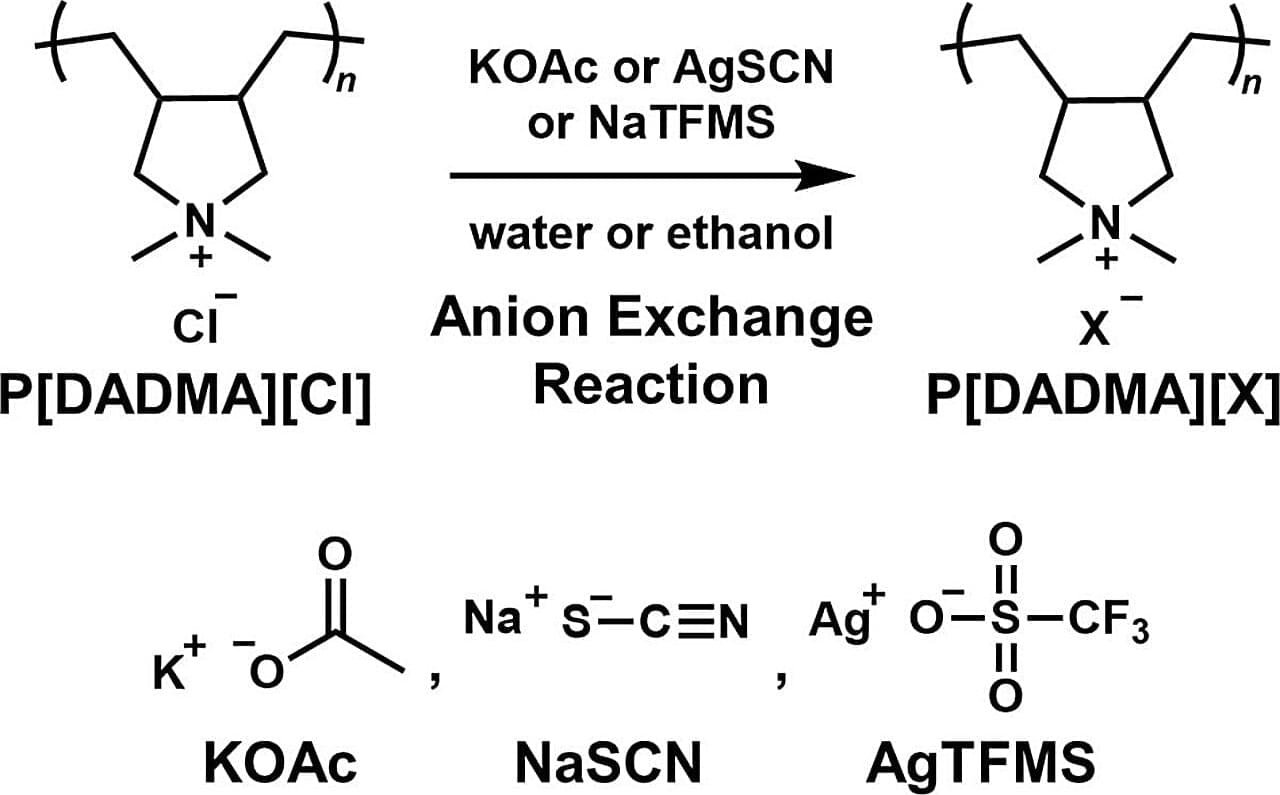
A joint research team from Nitto Boseki Co., Ltd. (Nittobo) and Tohoku University has revealed that polyionic liquids (PILs) can achieve high carbon dioxide (CO₂) adsorption when their counter anions are exchanged. This discovery provides a critical new design guideline for the development of high-performance CO2 recovery devices and gas separation membranes.
The research was led by Associate Professor Kouki Oka of the Institute of Multidisciplinary Research for Advanced Materials, Tohoku University, with the results published online in Reaction Chemistry & Engineering.
PILs are known for their strong ability to attract CO₂ and for their stability as solid materials. However, conventional anion exchange methods struggle to remove inorganic salts, which are by-products of the manufacturing process. These impurities make it difficult to accurately evaluate the materials’ true performance.
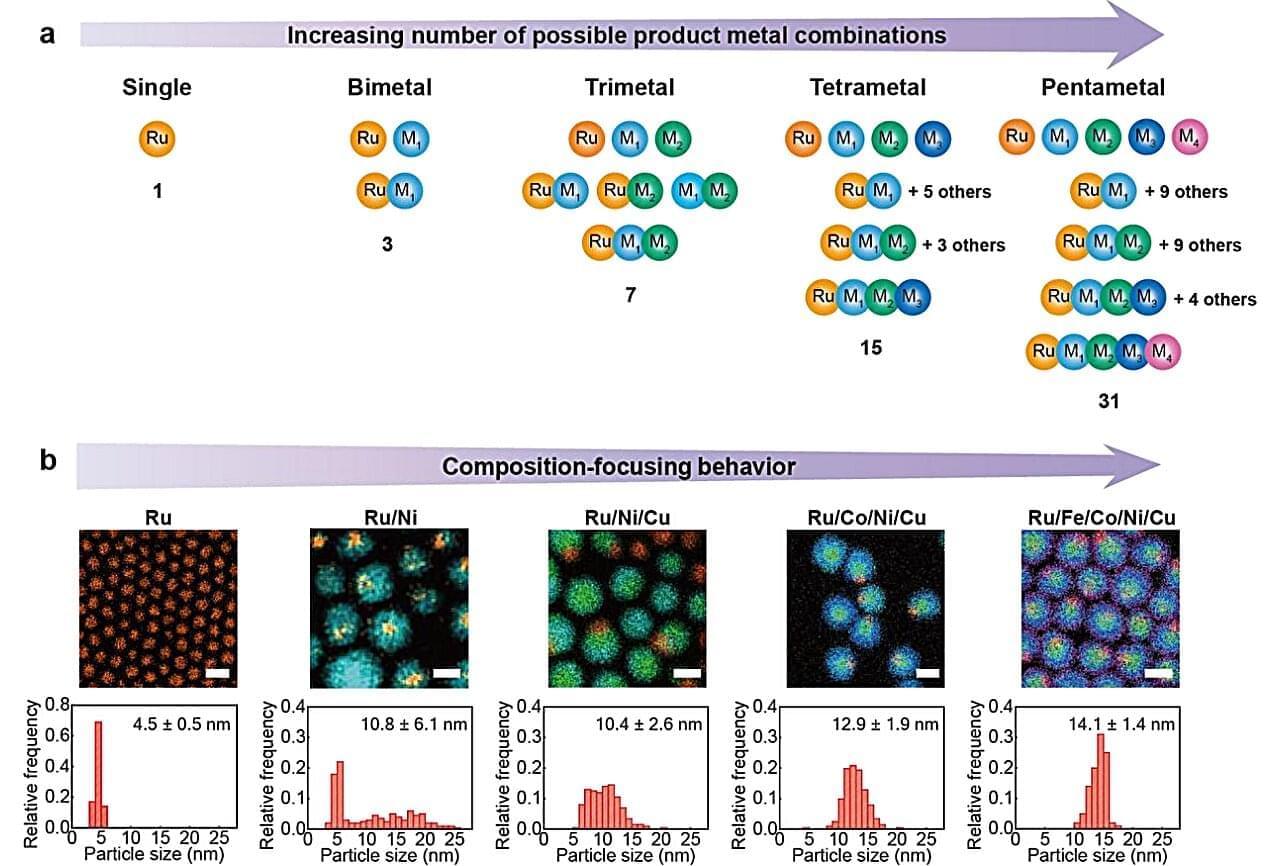
A nanocrystal is an extraordinarily tiny piece of material—composed of anywhere from a few to a few thousand atoms—in which atoms are arranged in a precise, ordered structure. Think of it like taking a piece of gold and shrinking it down to the size of a few hundred atoms. It’s still gold, still crystalline, just almost incomprehensibly small.
Nanocrystals are in the transistors inside computers and smartphones, in smartphone displays and TV screens, in the gold-nanoparticle sensors that power COVID and pregnancy tests, and in the pipes of your car exhaust system, among countless other innovations.
Their small size gives them a dramatically higher ratio of surface area to volume, making them especially useful as catalysts—materials that speed up chemical reactions without being consumed in the process.
Be kind to yourself this year. Using Zocdoc is FREE — visit my sponsor https://zocdoc.com/quinnsideas to find and instantly book an appointment with a top-rated, in-network doctor today.
Imagine a civilization reaches something like a Type II level, advanced enough to move through interstellar space and keep large populations alive for generations. At that stage, the challenge is developing ships that can cross the void, and also making sure the people inside them can survive radiation, isolation, and extreme travel times. That could mean heavy genetic engineering before the journey begins, changing bone density, metabolism, resistance to disease, tolerance for low gravity, or even sensory systems and respiration. But when they finally arrive, they may still find that the planet is wrong for them, maybe the air is toxic, the gravity is crushing, the temperatures are extreme, or the native chemistry is incompatible with human biology.
At that point, they face two paths. One is terraforming, which means trying to remake an entire planet into something closer to Earth. That could involve thickening or thinning an atmosphere, warming a frozen world, cooling a hot one, importing water, altering soil chemistry, introducing engineered microbes, building orbital mirrors or shades, and managing the planet for centuries or even millennia. The scale of that project is absurdly expensive, not just in money but in energy, infrastructure, labor, time, and raw materials. You are not changing a city or even a continent, you are trying to rewrite a whole world.
The other option is pantropy. Instead of forcing the planet to become Earth-like, the colonists change themselves to fit the planet. They might alter their lungs to breathe a different atmospheric mix, redesign their skin to handle harsher radiation, reduce their size for lower resource use, strengthen their bodies for higher gravity, or even become something so biologically different that they no longer look fully human. That is the core idea of pantropy, adapting the colonists to the world rather than adapting the world to the colonists.
The term was coined by James Blish, and he used it in connection with the stories collected in The Seedling Stars, especially “Surface Tension.” which was first published in 1952 in Galaxy Science Fiction.
Get 50% off Nebula (get access, early videos, exclusives): https://go.nebula.tv/quinnsideas.
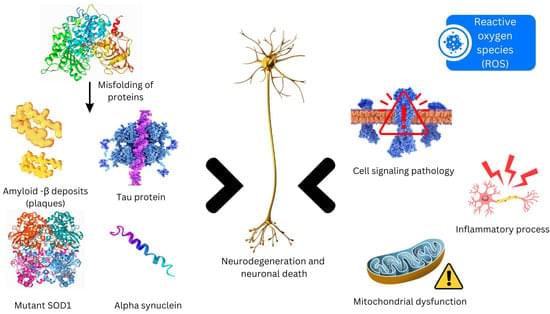
Neurodegenerative disorders entail a progressive loss of neurons in cerebral and peripheral tissues, coupled with the aggregation of proteins exhibiting altered physicochemical properties. Crucial to these conditions is the gradual degradation of the central nervous system, manifesting as impairments in mobility, aberrant behaviors, and cognitive deficits. Mechanisms such as proteotoxic stress, neuroinflammation, oxidative stress, and programmed cell death contribute to the ongoing dysfunction and demise of neurons. Presently, neurodegenerative diseases lack definitive cures, and available therapies primarily offer palliative relief. The integration of nanotechnology into medical practices has significantly augmented both treatment efficacy and diagnostic capabilities.

Researchers from the National University of Singapore (NUS) and collaborators have developed a predictive design strategy for creating graphene-like molecules with multiple interacting spins and enhanced resilience to magnetic perturbations, opening new avenues for molecular-scale quantum information technologies and next-generation spintronics.
The research team was led by Professor Lu Jiong from the NUS Department of Chemistry and the NUS Institute for Functional Intelligent Materials, together with Professor Wu Jishan from the NUS Department of Chemistry, and international collaborators, including key contributor Professor Pavel Jelínek from the Czech Academy of Sciences in Prague.
Magnetic nanographenes, which are molecules composed of fused benzene rings, are of growing interest for quantum technologies because they can host unpaired electrons, or spins, that may be used to store and process information. Unlike conventional magnetic materials based on metal atoms, these carbon-based systems offer chemical versatility and long spin coherence times. However, engineering a single molecule that contains multiple strongly coupled spins in a stable and controlled manner remains a major challenge.

Chemists and computer scientists tapped AI to find new disinfectants to combat the growing threat of dangerous “superbugs.”
The Journal of Chemical Information and Modeling published their computational-experimental framework for developing quaternary ammonium compounds, or QACs, to kill bacteria.
The method yielded 11 new QACs that show activity against antimicrobial-resistant bacteria.

Researchers at the University of Toronto have identified a protein from the quagga mussel that can stick to surfaces underwater, even though it lacks a chemical feature long thought to be essential for this kind of adhesion. The protein, called Dbfp7, is the first freshwater mussel adhesive protein to be functionally characterized.
The finding, published in PNAS, helps explain how some organisms attach themselves in wet environments and could inform the design of future medical glues—such as medical sealants and surgical adhesives—or other materials that need to work reliably in water.
Most studies of underwater adhesion have focused on marine mussels, which use proteins rich in a modified amino acid called 3,4-dihydroxyphenylalanine (DOPA) to bond to surfaces. Freshwater species have been studied less, and whether they rely on the same chemistry has not been clear.
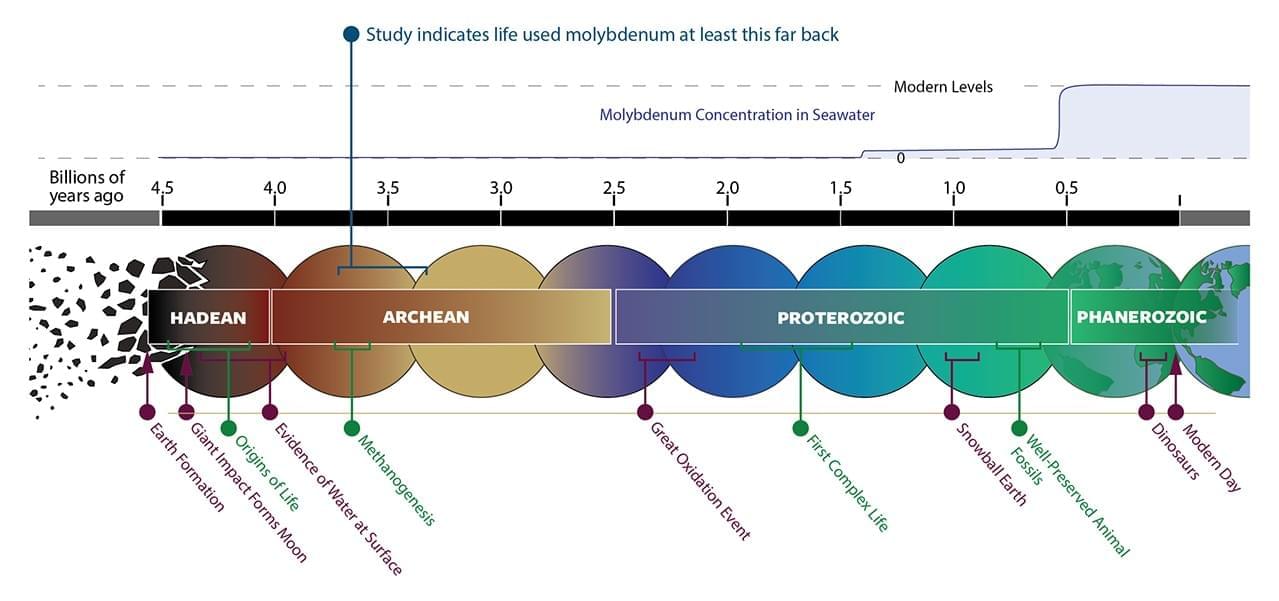
NASA-funded scientists have discovered that life on Earth over 3 billion years ago relied on the metal molybdenum, which was incredibly scarce in the environment at the time. The study, published in Nature Communications on Tuesday, is the first to show that molybdenum was used by ancient life this far back in our planet’s history.
On Earth today, molybdenum helps speed up vital biochemical reactions in cells. The metal is a component of essential enzymes that drive several major biological reactions in organisms. This is not only important for the individual organisms, but also biogeochemical cycles, such as the nitrogen cycle, which affect our entire planet. Without molybdenum, those important reactions could still happen in nature, but they would be too slow to sustain life.
“Molybdenum sits at the catalytic center of enzymes that run major carbon, nitrogen, and sulfur reactions,” explained Betül Kaçar, head of the Kaçar Lab at the University of Wisconsin-Madison and senior author on the study. Kaçar leads MUSE, a NASA Interdisciplinary Consortia for Astrobiology Research (ICAR) at UW-Madison.