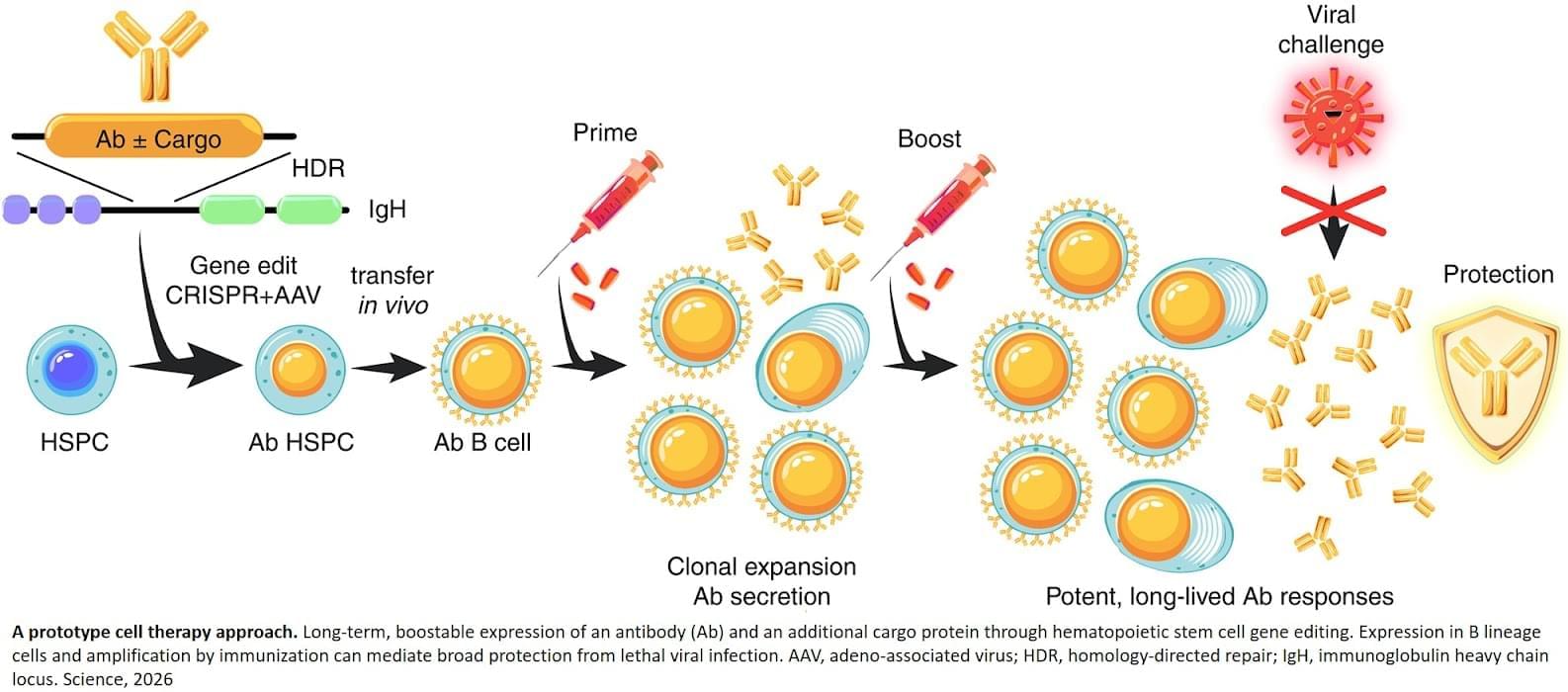
As a proof of concept, the team used CRISPR gene-editing tools to insert the genetic blueprint for producing rare, protective antibodies directly into hematopoietic stem and progenitor cells of mice. Once transplanted back into mice, the edited stem cells gave rise to B cells programmed to produce the engineered antibody. A conventional vaccination would then serve as the trigger.
It worked. Even when only a few dozen stem cells were edited, vaccination triggered rare cells to expand, mature into plasma cells, and produce large amounts of antibodies that persisted long-term and could be boosted if necessary. The engineered B cells behaved just like normal immune cells, and even provided protection from disease. Mice engineered to produce a broadly neutralizing influenza antibody were spared from an otherwise lethal influenza infection.
The team went on to demonstrate their novel platform’s versatility. Engineered B cells were able to secrete non-antibody proteins, pointing to potential applications in treating genetic diseases caused by missing enzymes or other essential proteins.
The researchers also showed that stem cells carrying different antibody instructions could be combined, enabling a single immune system to produce multiple antibodies at once—an approach that could limit viral escape and ultimately lead to functional cures for rapidly mutating pathogens such as HIV.
And the team showed that human stem cells edited using the same approach gave rise to functional immune cells, providing a key proof of feasibility that the platform could one day work in humans, as well. Science Mission sciencenewshighlights.
An innovative gene-editing strategy could establish a new way for the body to manufacture therapeutic proteins—including certain kinds of highly potent antibodies the are naturally difficult to produce—by reprogramming the immune system itself.