Researchers relied on a newer gene-editing technique that may make it possible to engineer embryos, a prospect that has long alarmed bioethicists.

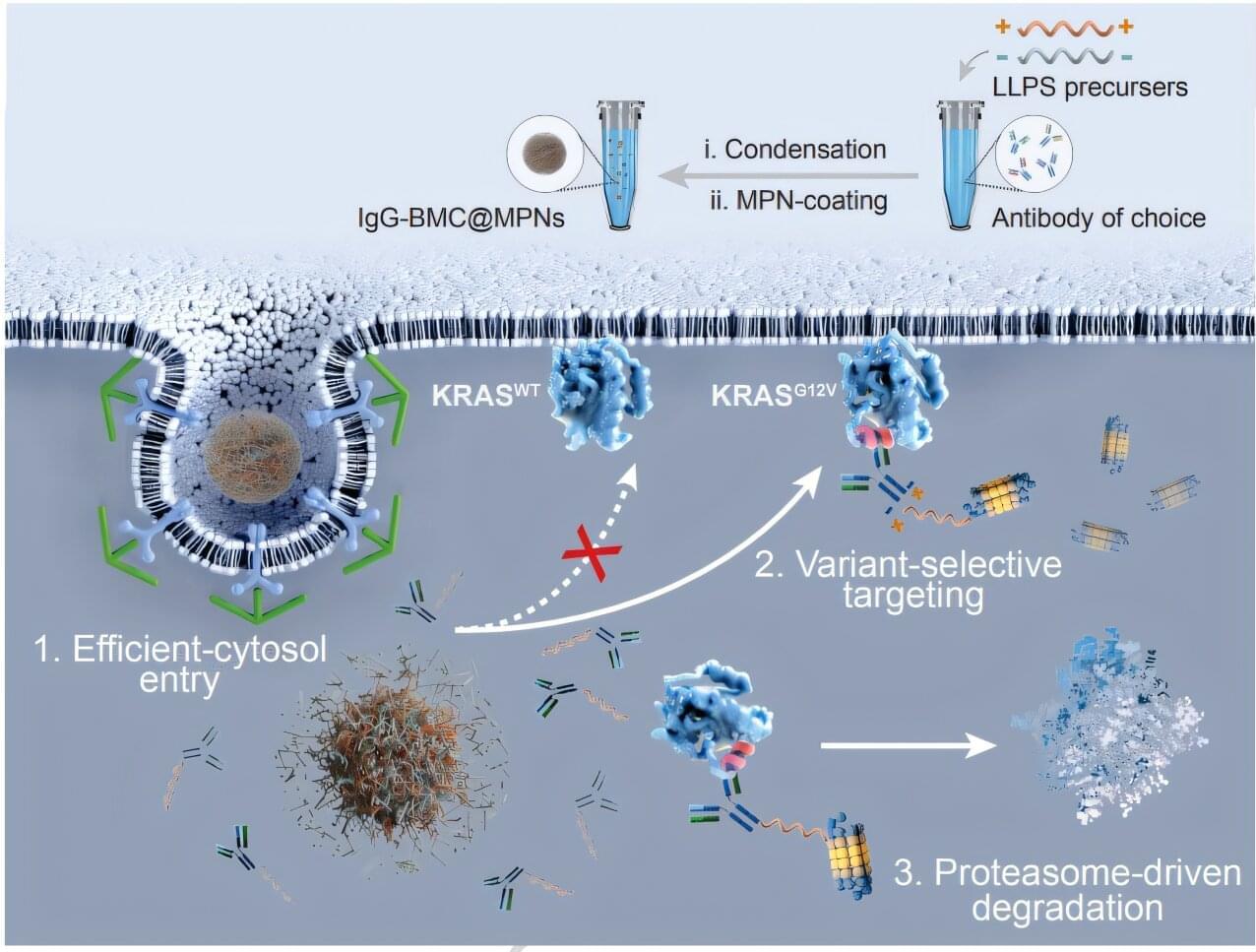
Northwestern Medicine scientists have developed a novel synthetic biomolecular condensate that can degrade intracellular disease-causing proteins, providing a framework for new therapeutic approaches for a wide range of diseases, as detailed in a recent study published in Nature Communications.
Shana Kelley, Ph.D., the Neena B. Schwartz Professor of Chemistry, Biomedical Engineering, and Biochemistry and Molecular Genetics and the president of the Chan Zuckerberg Biohub Chicago, was senior author of the study.
Targeted protein degradation is an emerging therapeutic strategy that harnesses cells’ own degradation machinery to clear disease-causing proteins. However, achieving this degradation process across different cell types has remained a challenge due to subtle variations in protein structure.

An international collaboration of genetic researchers has identified more than 90 genetic regions associated with the risk of Alzheimer’s disease and related dementias. The large-scale meta-analysis reveals new biological insights into the disease, highlighting the important roles of immune processes, beta-amyloid and tau biology, and lipid metabolism.
Alzheimer’s disease is the most common cause of dementia worldwide, and its development is influenced by a complex interplay of genetic and environmental factors. Understanding the genetic architecture of the disease is essential for improving diagnosis, risk prediction, and the development of targeted therapies.
In this study, researchers combined genome-wide association data from nearly a million individuals of European ancestry, including over 128,000 Alzheimer’s disease cases and nearly 850,000 controls.
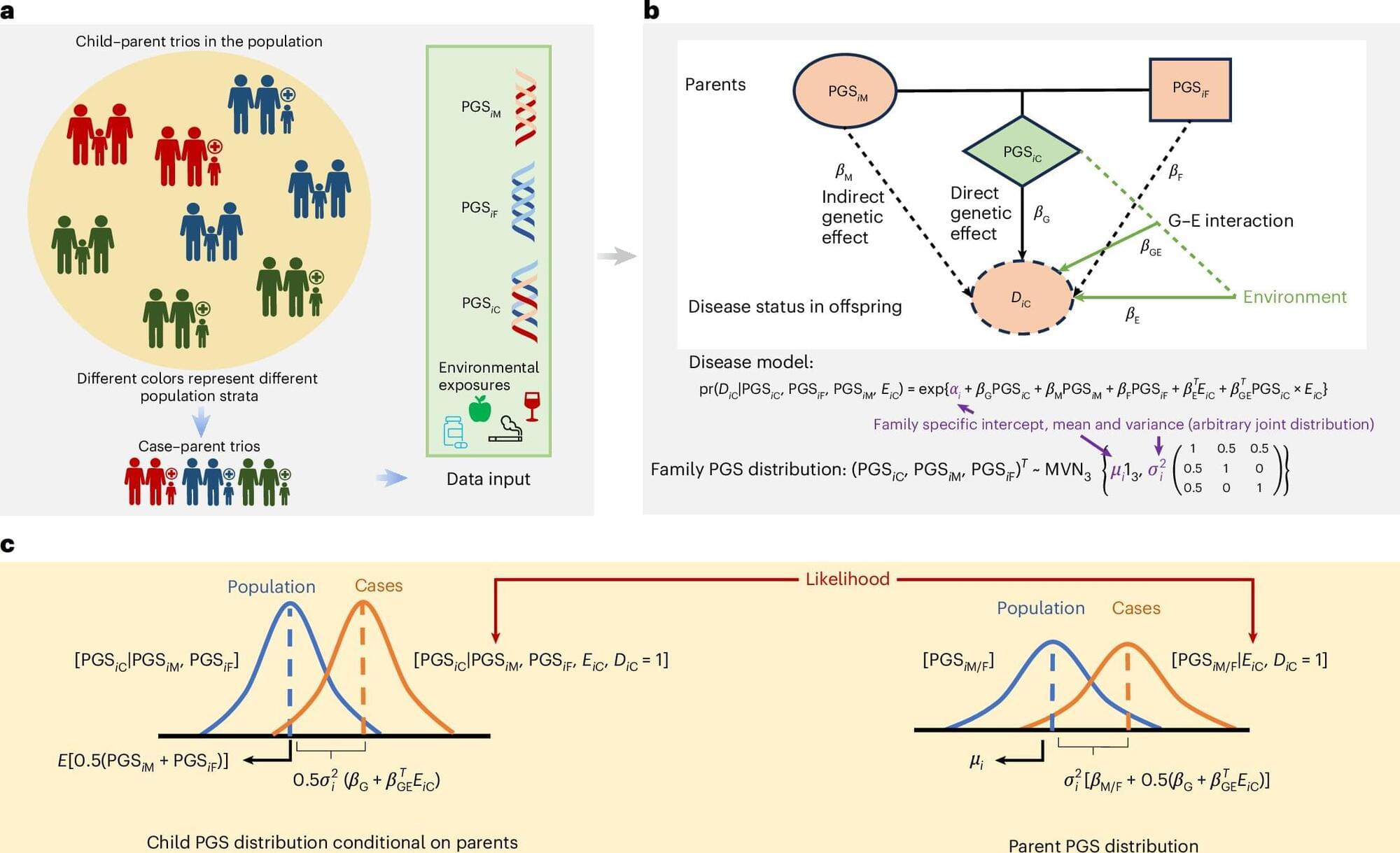
A new statistical framework developed by researchers at the Johns Hopkins Bloomberg School of Public Health, Johns Hopkins University School of Medicine, and Kaiser Permanente Northern California offers improved understanding of how genetics and environment contribute to autism risk.
Large-scale genetic studies have led to the development of genetic risk scores that estimate a person’s predisposition to diseases and health conditions based on their DNA profiles. The new framework allows researchers and clinicians to analyze these scores using family data and characterize the risk of conditions such as autism and other developmental conditions in children based on their own DNA, parental factors, and environmental influences such as maternal diet and lifestyle.
For their study published in Nature Genetics, the researchers analyzed more than 18,000 case-parent trios —autistic children and their parents—across diverse ancestral populations in the Simons Foundation Powering Autism Research for Knowledge consortium and the Genes and Environment Autism Research Study.
On April 21, the Munk Debates convened a special debate about gene editing in Deerfield, Massachusetts for 650 students at Deerfield Academy.
Motion: Be it Resolved, let’s engineer better human beings.
About the Debate:
New powerful engineering technology is already being used to edit human embryos, curing diseases and repairing defective genes before a child is even born. Some welcome this new science as a powerful tool to enhance human intelligence, memory, appearance and physical health. Why wouldn’t we embrace a science that allows people to live longer, healthier, and happier lives? Others warn that this new technology will be used to create designer babies and a new class of genetically “enhanced” elites. It will undermine human dignity and autonomy, and risk unleashing new diseases into the human gene pool. Playing G-d with human nature, critics argue, will result in a dystopian nightmare of our own making.
About the Debaters:
Arguing in favour of the motion was the biophysicist, best-selling author, biotechnology entrepreneur, and the former director of the Program on Medicine, Technology and Society at UCLA School of Medicine, Gregory Stock. His debate partner was the internationally acclaimed strategic philosopher and pioneering transhumanist Max More. Arguing against the motion was the prominent American bioethicist Ezekiel Emanuel, Special Advisor to the Director General of the WHO and a former founding chair of the Department of Bioethics at the NIH. His debate partner was the award-winning educator, author, and Professor of Reproductive Science at University College London, Joyce Harper.
In this groundbreaking conversation, Professor of Genetics and longevity scientist, Dr. David Sinclair, A.O., Ph.D., joins Sarah Grynberg to unpack the future of human aging, the science of longevity, and how we live today impacts how we age tomorrow.
From reversing blindness in mice to exploring treatments that could one day delay menopause and extend healthy human life, this episode will completely change the way you think about your body, your health, and your future.
But beyond the science, this is also a deeply human conversation about purpose, suffering, love, family, and what it truly means to live a great life.
In this episode, you will learn:
Why aging may actually be reversible.
The daily habits accelerating aging in your body right now.
How stress, loneliness, and cortisol could impact longevity.
The real science behind supplements like NMN, resveratrol, and NAD boosters.
Why exercise, sleep, and relationships matter more than you think.
What Dr. Sinclair believes is coming in the next 10 years of medicine.
How scientists are working to reverse female infertility and delay menopause.
The surprising reason your “biological age” may be younger or older than your real age.
Why suffering through disease and decline should not be considered “normal aging”
The philosophy and mindset Dr. Sinclair lives by every day.
00:00 — Introduction.
01:18 — Why David Sinclair Became Obsessed With Aging.
06:20 — The Childhood Conversation That Changed His Life.
10:18 — The Groundbreaking Discovery That Could Reverse Aging.
12:47 — Reversing Blindness In Mice.
13:33 — Human Trials Are About To Begin.
16:11 — What Accelerates Aging Faster Than Anything Else.
20:08 — Why Relationships & Loneliness Impact Longevity.
24:14 — The Truth About Sun Exposure & Aging.
28:59 — Alzheimer’s, Cancer & Diseases Of Aging.
35:28 — Will Humans Live Longer In The Next Decade?
38:34 — The Supplements David Sinclair Personally Takes.
46:50 — Menopause, Fertility & Reversing Ovarian Aging.
50:20 — What Humans Will Eventually Die From.
51:18 — The Difference Between His Mother & Father’s Aging.
55:37 — Skin Rejuvenation, Hair Growth & Looking Younger.
58:16 — Why He Became A “Struggling Vegan”
01:00:08 — David Sinclair’s Workout & Exercise Routine.
01:03:28 — The Lifespan Community & Podcast.
01:06:02 — The Best Advice He’s Ever Received.
01:08:09 — What A Life Of Greatness Means To David Sinclair.
This episode is a powerful reminder that longevity is not just about living longer… it’s about living better.
Hookworms, intestinal parasites that infect hundreds of millions of people in under-resourced tropical regions around the globe, have evolved to survive inside the human gut for years, secreting molecules that enable coexistence with their hosts. Now, researchers at Washington University School of Medicine in St. Louis have harnessed that biological mechanism for potential human benefit, engineering a hookworm to produce and deliver a drug within a living host.
In a new study, the team reports the first successful genetic modification of the human hookworm. It was designed to produce an antibody that neutralizes tetrodotoxin, a deadly neurotoxin produced by pufferfish and other marine animals. After colonizing an animal host with the modified hookworms, the parasites produced the antitoxin and secreted it into the bloodstream, partially inactivating the toxin. The findings are published in Nature Communications.
The work demonstrates that this drug production and delivery approach could be a long-term solution to any number of medical needs, from chronic conditions requiring continuous drug treatment to exposure to toxins in remote locations without medical care available.

A Phase III clinical trial led by Neeraj Agarwal, MD, FASCO, senior director of clinical research at Huntsman Cancer Institute and professor of internal medicine at the University of Utah (the U), has found that a combination prostate cancer treatment could prevent the disease from progressing into a harder-to-treat form of cancer in select patients.
The study, TALAPRO-3 (NCT04821622), evaluated a combination of two drugs—talazoparib and enzalutamide—in patients with metastatic castration-sensitive prostate cancer. This is a form of the disease that has spread beyond the prostate but remains susceptible to standard hormone therapy treatment.
The patients involved also had prostate cancer affected by certain gene mutations, including but not limited to BRCA1 and BRCA2 mutations, that often signal more aggressive disease.
Abstract Despite substantial progress in identifying neural correlates of consciousness, no unified quantitative framework currently derives a formally specified order parameter for conscious-state organisation from established neurophysiological principles, or links thalamocortical coordination dynamics to measurable state transitions across pharmacological, pathological, and perturbational conditions through a single computational formalism. We propose a neurocomputational theoretical framework in which conscious states are associated with metastable regimes of large-scale thalamocortical coordination operating near critical dynamical boundaries. The framework is formalised through a dynamic coordination functional Φ(t), defined as a surface integral over the thalamocortical interface and directly operationalisable from high-density EEG as a weighted combination of gamma-band power spectral density, thalamocortical coherence, and theta-gamma phase-amplitude coupling. The thalamic reticular nucleus (TRN) is identified as the anatomical implementation of the control parameter governing proximity to the critical point, grounded in a Wilson-Cowan model of TRN inhibitory gating whose bifurcation structure is characterised computationally. Numerical simulation of the linearised field equation on the thalamocortical boundary demonstrates internal consistency: the simulated system produces power-law recovery dynamics tau_rec proportional to | θ — θ _c|^v with nu consistent with model A universality class [0.5, 1.5], and a Kuramoto mean-field derivation establishes that Φ(t) emerges as the natural order parameter of coupled thalamocortical oscillators rather than being postulated. The joint (|Φ(t)|, Var[|Φ(t)|]) phase space correctly separates simulated waking, anaesthetic, ictal, and minimally conscious regimes without parameter fitting to empirical data. All simulation code is publicly available. Six quantitatively specific, independently falsifiable predictions are derived across five experimental domains: power-law Gamma Dip scaling in near-threshold EEG with a specific exponent range; causal disruption of thalamocortical coherence by selective TRN silencing; opposite EEG scaling exponent deviations in ASD versus schizophrenia; systematic Φ_est collapse under propofol anaesthesia correlated with PCI; Φ_est as a real-time consciousness biomarker in disorders of consciousness; and clinical validity of Φ_est in disorders of consciousness and ictal state discrimination by the metastability index. Each prediction is stated with quantitative thresholds and a pre-specified falsification criterion. The framework provides: the first anatomically specified and formally derived order parameter for conscious-state organisation directly operationalisable from passive EEG; a mechanistically grounded identification of the TRN as the dynamical control parameter, testable by a single optogenetic experiment; and a computationally validated, pre-registerable programme of six falsifiable predictions defining a tractable empirical agenda. Φ_est would constitute a candidate real-time consciousness biomarker if the framework’s predictions are confirmed in purpose-designed experiments.
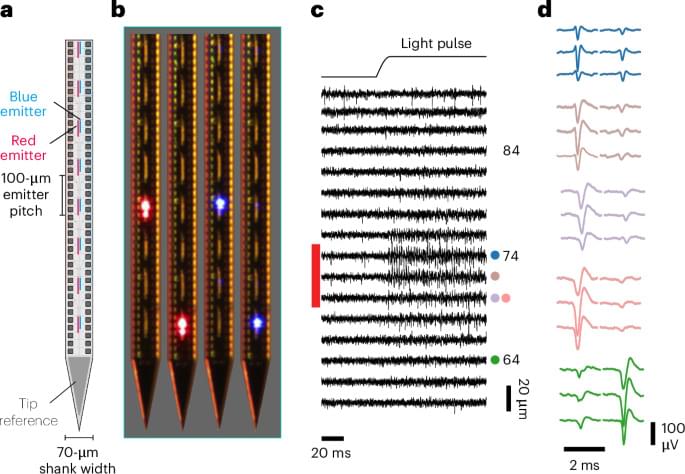
High-density electrophysiology devices allow neuroscientists to observe spikes from large populations of neurons, and optogenetics allows them to drive or suppress those spikes. We show that a single device can combine these two capabilities, providing a high-resolution means to both read and write neural activity in the living brain.