ITrust Capital: Use code IMPACTGO when you sign up and fund your account to get a $100 bonus at https://impacttheory.co/iTrustCapitalMay.
Shopify: Sign up for your one-dollar-per-month trial period at https://impacttheory.co/ShopifyApr.
On this mind-bending episode of Impact Theory, Tom Bilyeu sits down with Ben Lamm, the visionary entrepreneur behind Colossal Biosciences, to explore a world that sounds straight out of science fiction—yet is rapidly becoming our reality. Together, they pull back the curtain on the groundbreaking technology making de-extinction not only possible, but increasingly practical, from resurrecting woolly mammoths and dire wolves to saving endangered species and unraveling the secrets of longevity.
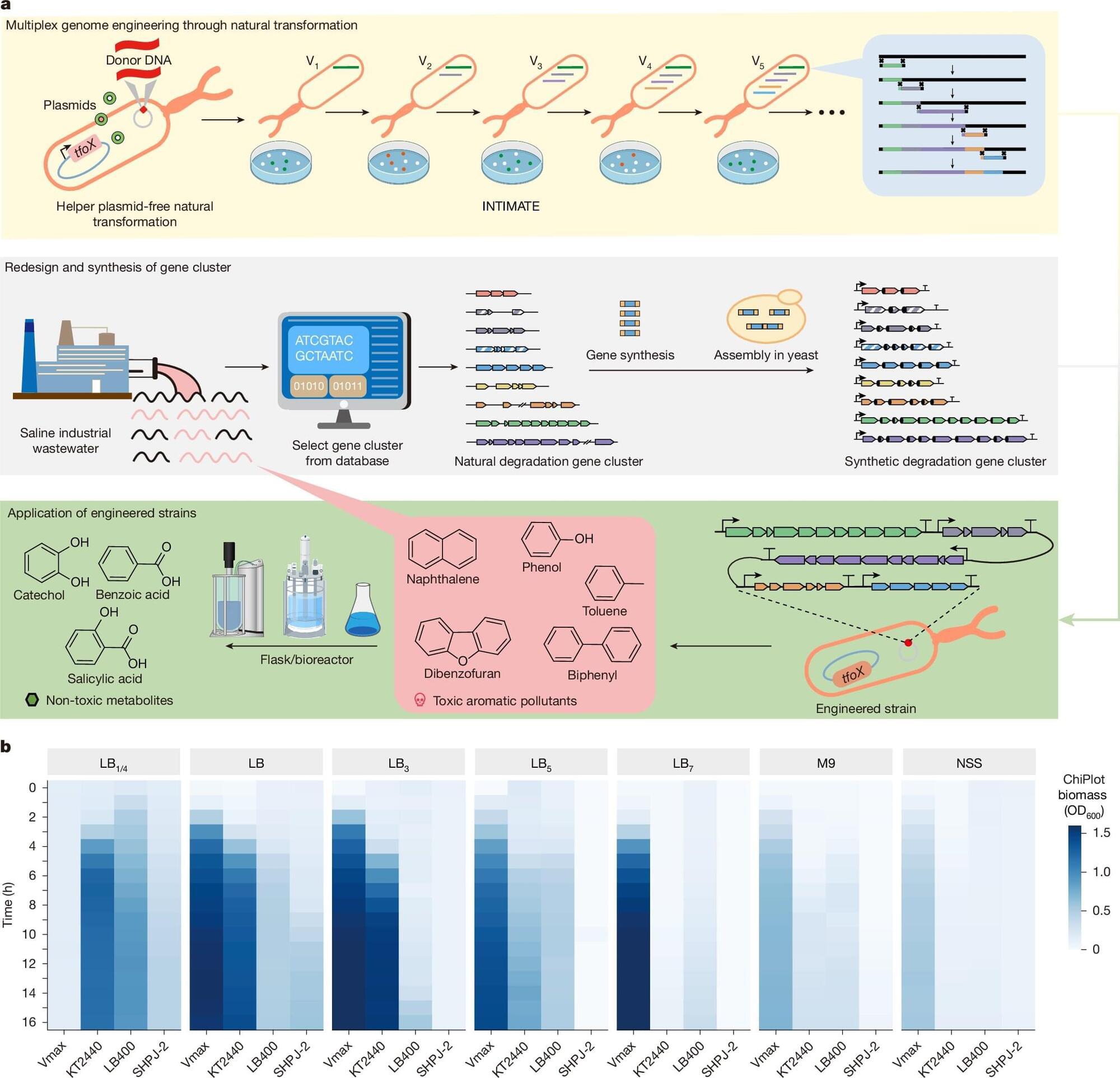
Ben explains how CRISPR gene editing has unlocked the power to make precise DNA changes—editing multiple genes simultaneously, synthesizing entirely new genetic blocks, and pushing the limits of what’s possible in biology and conservation. The conversation dives deep into the technical hurdles, ethical questions, and the unexpected magic of re-engineering life itself, whether it’s creating hairier, “woolly” mice or tackling the colossal challenge of artificial wombs and universal eggs.
But this episode goes way beyond Jurassic Park fantasies. Tom and Ben debate the future of human health, gene selection through IVF, the specter of eugenics, global competition in biotechnology, and how AI will soon supercharge the pace of biological engineering. They even touch on revolutionary solutions to our plastic crisis and what it means to inspire the next generation of scientists.
Get ready to have your mind expanded. This is not just a podcast about bringing back extinct creatures—it’s a deep dive into the next frontiers of life on Earth, the technologies changing everything, and the choices we’ll face as architects of our own biology. Let’s get legendary.
00:00 Meet Ben Lamm.