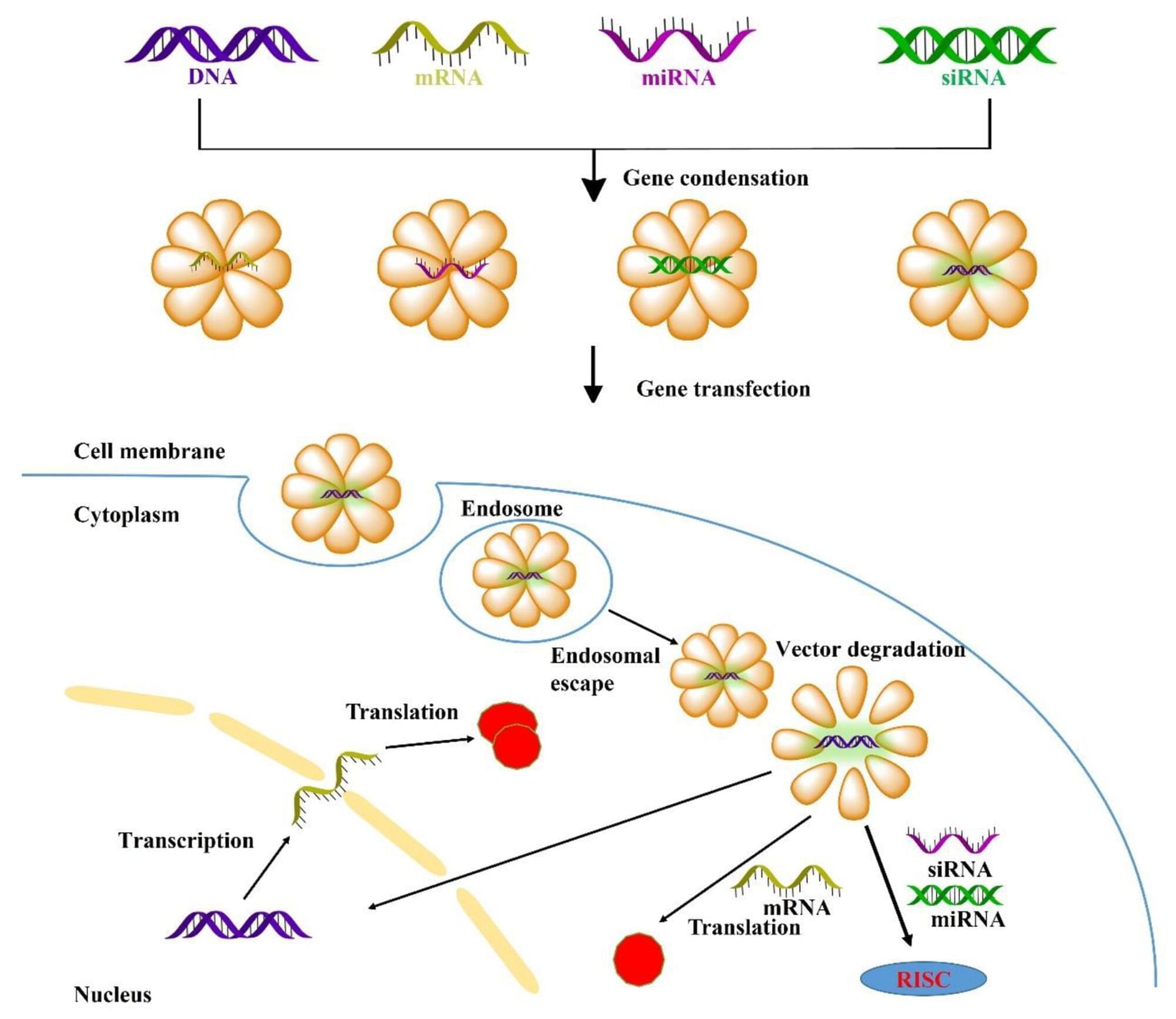
The development of COVID-19 vaccines has sparked widespread interest. mRNA-based therapies are rapidly gaining attention owing to their unique advantages in quickly developing vaccines and immunotherapy for various ailments [1, 2]. Given that most human diseases stem from genetic factors, gene therapy represents a promising modality for addressing various inherited or acquired disorders by replacing faulty genes or silencing genes [3]. Gene therapy encompasses the targeted exploitation of genetic material, which includes gene replacement through DNA or mRNA [4, 5]; gene silencing utilizing siRNA or miRNA [6], and CRISPR-Cas9 based gene editing [7].
However, achieving safe and efficient gene delivery to specific cells requires overcoming multiple biological barriers, including extracellular obstacles such as enzyme degradation, serum protein interactions, electrostatic repulsion of genes and cell membranes, and innate immune system, as well as intracellular obstacles such as endosomal escape, transport barriers, precise release [8]. Therefore, gene vectors require several characteristics such as high gene condensation; favorable serum stability to avoid non-specific serum protein interactions, endonuclease degradation, and renal clearance; achieved specific targeting cell or tissues; effective transport into the cytoplasm thereby facilitating gene transfection (mRNA, siRNA and miRNA); precise gene release and scheduling, and nuclear localization that enables DNA transcription. Comprehensive exploration of transfection mechanisms can aid in the development of high-performance gene vectors [9, 10].
Gene vectors generally include viral vectors and non-viral vectors. Presently, approximately 70% of clinical gene therapy trials employ viral vectors, which include retroviruses, lentiviruses, adenoviruses, and adeno-associated viruses. Due to their exceptional infectivity, virus-based vectors typically exhibit excellent gene transfection capabilities. However, the clinical safety of viral vectors has been questioned due to their propensity to stimulate immunogenic reactions and induce transgene insertion mutations. Moreover, viral vectors possess several limitations, including low gene loading capacity, inability to deliver large-sized genes, complicated preparation procedures, and the patient cannot be repeatedly administered [4]. In contrast, non-viral vectors, particularly lipid nanoparticles (LNPs) and cationic polymers, have demonstrated robust gene loading capacity, heigh safety and practicability, simplicity preparation [10, 11]. Consequently, non-viral vectors are exhibiting tremendous potential for further clinical development and application. Our review primarily highlights the significant potential of non-viral vectors, particularly lipid nanoparticles (LNPs), highly branched poly(β-amino ester) (HPAE), single-chain cyclic polymer (SCKP), poly(amidoamine) (PAMAM) dendrimers, and polyethyleneimine (PEI). We intend to provide a detailed examination of the latest research progress and existing limitations of non-viral gene vectors over recent years.