We are used to heat flowing from hot objects to cool ones, and never the other way round, but now researchers have found it is possible to pull off this trick in the strange realm of quantum mechanics
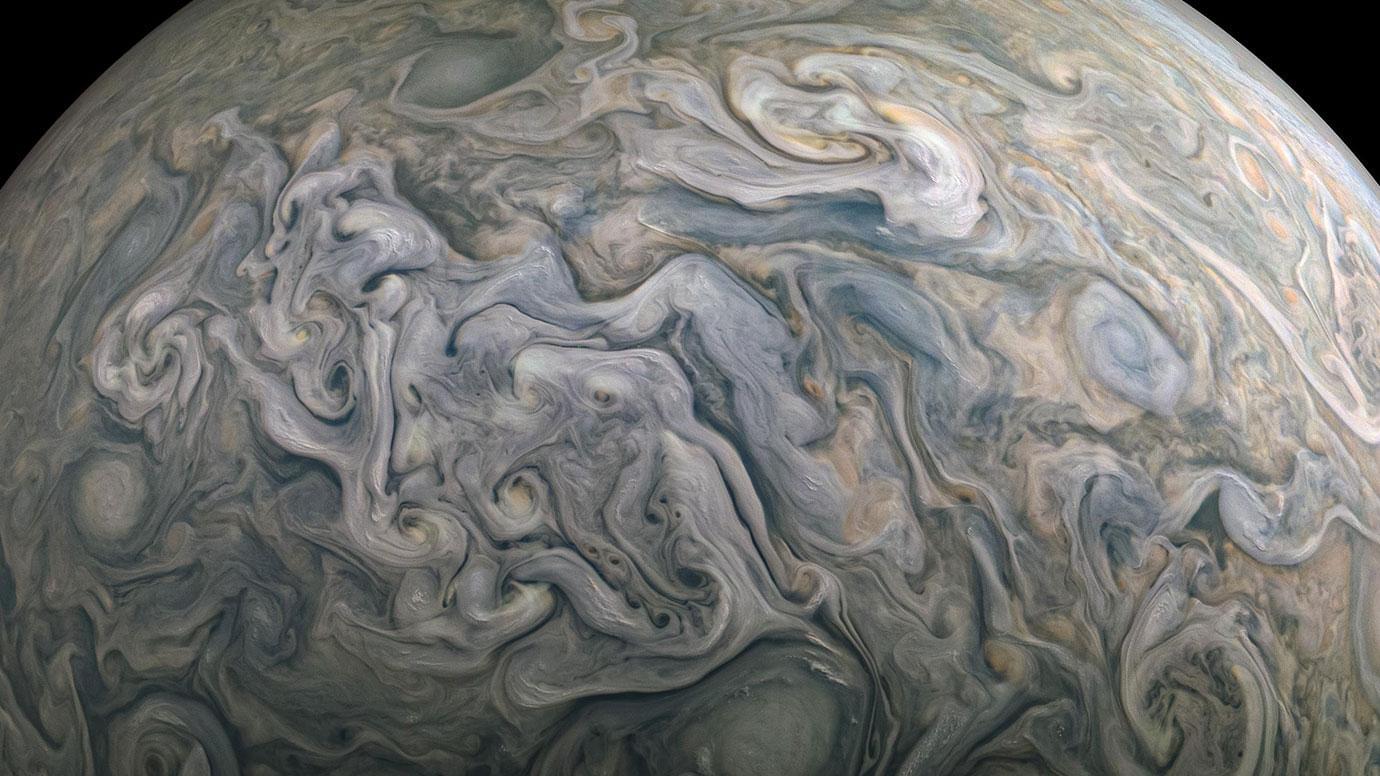






Jupiter’s swirling storms have concealed its true makeup for centuries, but a new model is finally peeling back the clouds. Researchers found the planet likely holds significantly more oxygen than the Sun, a key clue to how Jupiter—and the rest of the solar system—came together. The study also reveals that gases move through Jupiter’s atmosphere much more slowly than scientists once thought. Together, the findings reshape our understanding of the solar system’s largest planet.
Towering clouds ripple across Jupiter’s surface in dramatic patterns. Like Earth’s clouds, they contain water, but on Jupiter they are far denser and far deeper. These layers are so thick that no spacecraft has been able to directly observe what lies below them.
Now, scientists have taken a major step toward solving that mystery. A new study led by researchers at the University of Chicago and the Jet Propulsion Laboratory has produced the most detailed model of Jupiter’s atmosphere ever created. The work provides a deeper look into the planet’s interior without needing to physically descend into its crushing depths.

A company called Fractal Health is developing a one-time GLP-1 gene therapy and the first humans are being dosed this year.
Instead of weekly injections like semaglutide or tirzepatide, this approach delivers genes directly into pancreatic beta cells. The therapy is controlled by the insulin promoter, meaning GLP-1 is only released when you eat — not continuously.
That could mean fewer systemic side effects and a more physiologic response.
They’re developing two versions:
• GLP-1 alone.
• GLP-1 + GIP (similar to tirzepatide)
One injection. Potentially permanent metabolic support.
If this works, it could redefine obesity and diabetes treatment.
Industrial yeasts are a powerhouse of protein production, used to manufacture vaccines, biopharmaceuticals, and other useful compounds. In a new study, MIT chemical engineers have harnessed artificial intelligence to optimize the development of new protein manufacturing processes, which could reduce the overall costs of developing and manufacturing these drugs.
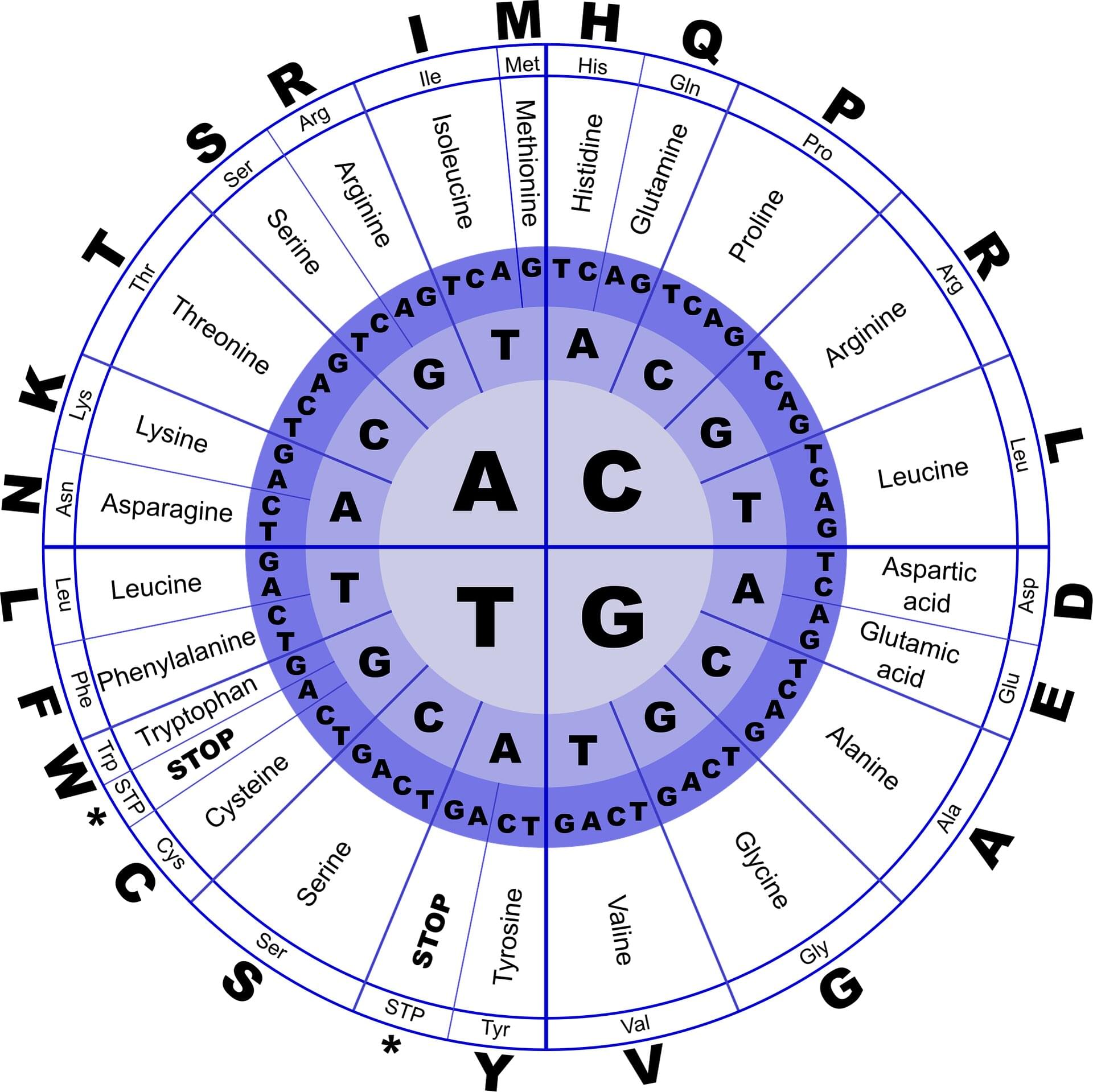
Using a large language model (LLM), the MIT team analyzed the genetic code of the industrial yeast Komagataella phaffii—specifically, the codons that it uses. There are multiple possible codons, or three-letter DNA sequences, that can be used to encode a particular amino acid, and the patterns of codon usage are different for every organism.
The new MIT model learned those patterns for K. phaffii and then used them to predict which codons would work best for manufacturing a given protein. This allowed the researchers to boost the efficiency of the yeast’s production of six different proteins, including human growth hormone and a monoclonal antibody used to treat cancer.