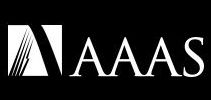There’s some really interesting CRISPR news out today, and it’s likely to be a forerunner of much more news to come. A research team has demonstrated what looks like robust, long-lasting effects in a primate model after one injection of the CRISPR enzymatic machinery. There have been plenty of rodent reports on various forms of CRISPR, and there are some human trials underway, but these is the first primate numbers that I’m aware of.
The gene they chose to inactivate is PCSK9, which has been a hot topic in drug discovery for some years now. It’s a target validated by several converging lines of evidence from the human population (see the “History” section of that first link). People with overactive PCSK9 have high LDL lipoproteins and cholesterol, and people with mutations that make it inactive have extremely low LDL and seem to be protected from a lot of cardiovascular disease. There are several drugs and drug candidates out there targeting the protein, as well there might be.
It’s a good proof-of-concept, then, because we know exactly what the effects of turning down the expression of active PCSK9 should look like. It’s also got the major advantage of being mostly a liver target – as I’ve mentioned several times on the blog already, many therapies aimed at gene editing or RNA manipulation have a pharmacokinetic complication. The formulations used to get such agents intact into the body (and in a form that they can penetrate cells) tend to get combed out pretty thoroughly by the liver – which after all, is (among other things) in the business of policing the bloodstream for weird, unrecognized stuff that is then targeted for demolition by hepatocytes. Your entire bloodstream goes sluicing through the liver constantly; you’re not going to able to dodge it if your therapy is out there in the circulation. It happens to our small-molecule drugs all the time: hepatic “first pass” metabolism is almost always a factor to reckon with.