Combining a map of gene regulatory sites with disease-associated loci has uncovered a new genetic risk factor of adult-onset macular degeneration (AMD), according to a new study publishing January 17 in the open access journal PLOS Biology by Ran Elkon and Ruth Ashery-Padan of Tel Aviv University, Israel, and colleagues. The finding advances the understanding of the leading cause of visual impairment in adults.
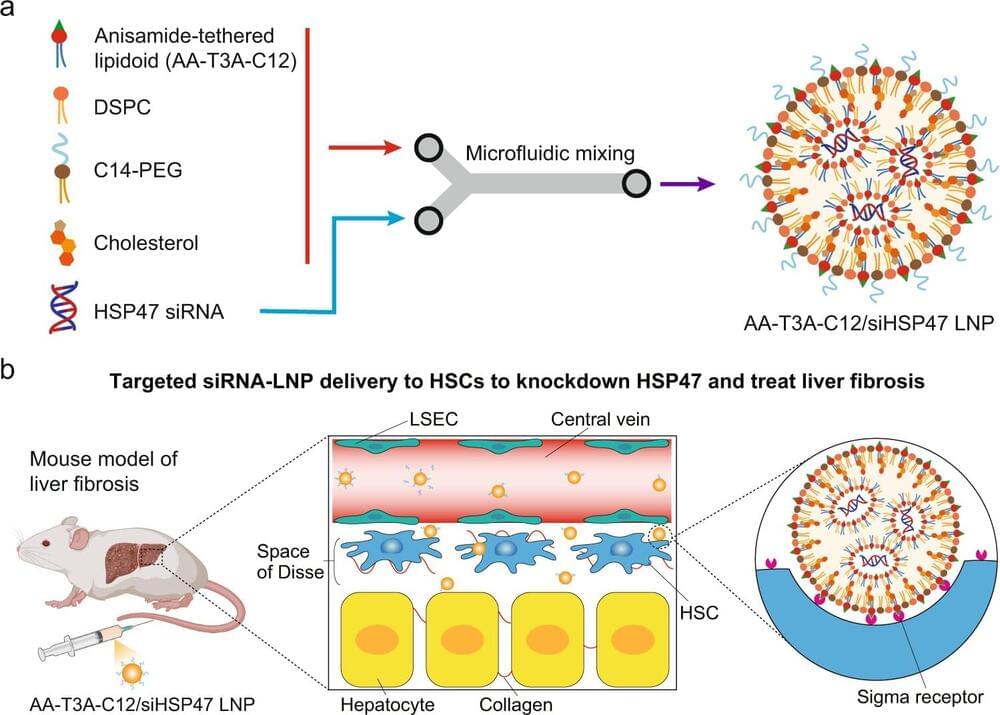
AMD is caused by dysfunction in the retinal pigmented epithelium (RPE), a layer of tissue sandwiched between the photoreceptors that receive light, and the choriocapillaris, which nourishes the retina. Because of the central importance of the RPE in AMD, the authors began by exploring a transcription factor (a protein that regulates specific genes) called LHX2 which, based on the team’s analysis of mouse mutants, is central to RPE development. Knocking down LHX2 activity in RPE derived from human stem cells, they found that most affected genes were down-regulated, indicating that LHX2’s role was likely that of a transcriptional activator, binding to regulatory sites on the genome to increase activity of other genes.
The authors found that one affected gene, called OTX2, collaborated with LHX2 to regulate many genes in the RPE. By mapping the genomic sites that OTX2 and LHX2 could bind to, they showed that 68% of those that bound LHX2 were also bound by OTX2 (864 sites in all), suggesting they likely work together to promote the activity of a large suite of genes involved in RPE development and function.