How we’re linking together genetic material from thousands of people — modern and ancient — to trace our ancestors and the history of our evolution.



Summary: A novel study uncovers a peculiar pattern of decision-making in mice, influenced by a specific gene named Arc.
While searching for food, mice repeatedly visited an empty location instead of staying at a site abundant in food. However, mice lacking the Arc gene demonstrated a more practical approach, sticking with the food-rich site, thereby consuming more calories overall.
This unique research potentially opens the door for a new field, ‘decision genetics’, investigating the genetic influence on decision-making, possibly even in humans.
Join us on Patreon! https://www.patreon.com/MichaelLustgartenPhD
Discount Links:
At-Home Metabolomics: https://www.iollo.com/?ref=michael-lustgarten.
NAD+ Quantification: https://www.jinfiniti.com/intracellular-nad-test/
Use Code: ConquerAging At Checkout.
Green Tea: https://www.ochaandco.com/?ref=conqueraging.
Oral Microbiome: https://www.bristlehealth.com/?ref=michaellustgarten.
Enter Code: ConquerAging.
Epigenetic Testing: https://trudiagnostic.com/?irclickid=U-s3Ii2r7xyIU-LSYLyQdQ6…M0&irgwc=1
Scientists have enhanced the efficiency of CRISPR/Cas9 gene editing by threefold using interstrand crosslinks, without resorting to viral material for delivery. This approach boosts the cell’s natural repair mechanisms, allowing for more accurate and efficient gene editing, potentially improving disease research and preclinical work.
Gene editing is a powerful method for both research and therapy. Since the advent of the Nobel Prize-winning CRISPR/Cas9 technology, a quick and accurate tool for genome editing discovered in 2012, scientists have been working to explore its capabilities and boost its performance.
Researchers in the University of California, Santa Barbara biologist Chris Richardson’s lab have added to that growing toolbox, with a method that increases the efficiency of CRISPR/Cas9 editing without the use of viral material to deliver the genetic template used to edit the target genetic sequence. According to their new paper published in the journal Nature Biotechnology, their method stimulates homology-directed repair (a step in the gene editing process) by approximately threefold “without increasing mutation frequencies or altering end-joining repair outcomes.”
I quoted and responded to this remark:
“…we probably will not solve death and this actually shouldn’t be our goal.” Well nice as she seems thank goods Dr Levine does not run the scientific community involved in rejuvenation.
The first bridge looks like it’s going to be plasma dilution and this may come to the general population in just a few short years. People who have taken this treatment report things like their arthritis and back pain vanishing.
After that epigentic programming to treat things that kill you in old age. And so on, bridge after bridge. if you have issues with the future, some problem with people living as long as they like, then by all means you have to freedom to grow old and die. That sounds mean but then I think it’s it’s mean to inform me I have to die because you think we have to because of “progress”. But this idea that living for centuries or longer is some horrible moral crime just holds no water.
Science can’t stop aging, but it may be able to slow our epigenetic clocks.
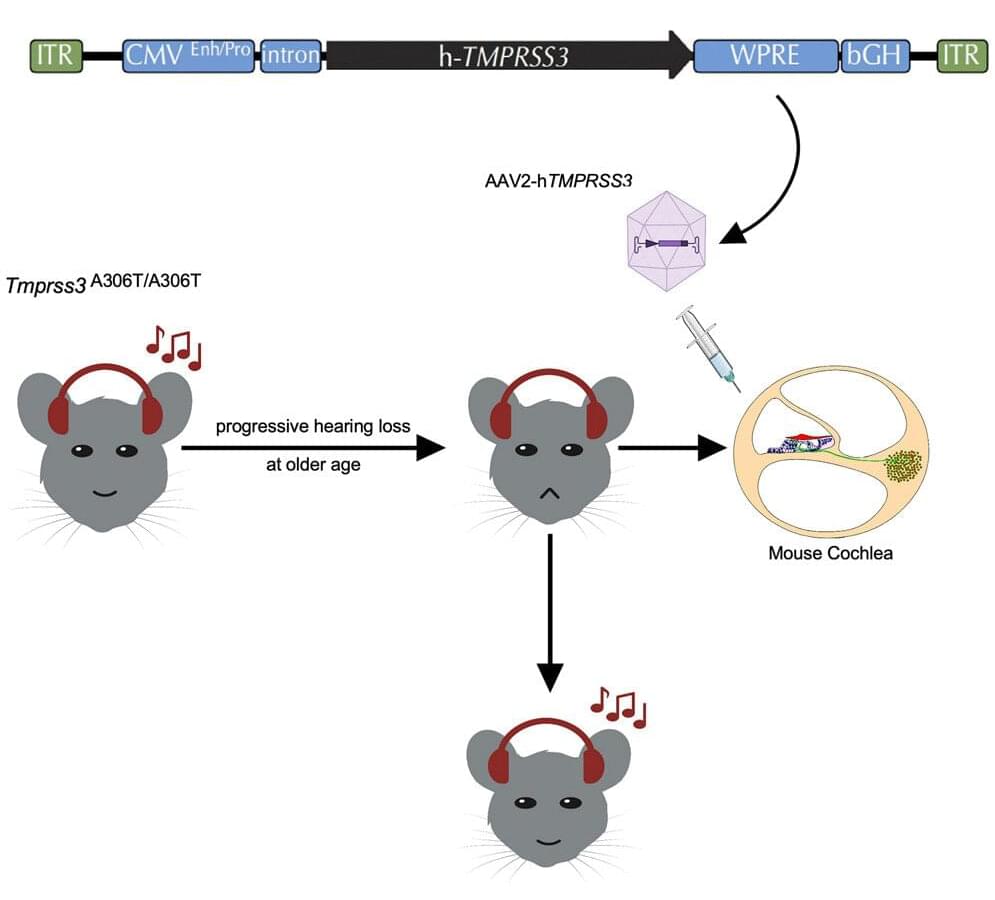
While hearing aids and cochlear implants offer limited relief, no available treatment can reverse or prevent this group of genetic conditions, prompting scientists to evaluate gene therapies for alternative solutions.
One of the most promising tools used in these therapies—adeno associated virus (AAV) vectors—has galvanized the hearing-loss community in recent years.
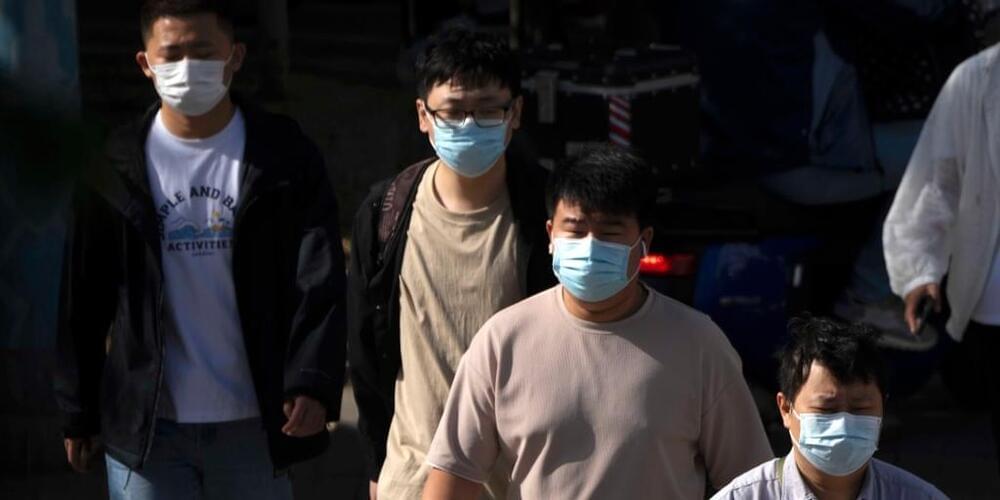

So it is confirmed that the new variant of covid 19 virus is here but the actual spike now is in China. But will most likely spread globally much how previous viruses have done. Be sure to be prepared for another pandemic. Anyway what may be the possible cure would be new bioengineering techniques with crispr to eventually be immune to the virus like I have posted in some genetically engineered cells recently were made. But rest assured this could lead to a global pandemic because the current variant is taxing our current vaccination measures.
The country once had some of the harshest Covid restrictions on the planet, but the response from the government and the public is relatively muted this time.
With the new method, scientists can explore many cancer mutations whose roles are unknown, helping them develop new drugs that target those mutations.
MIT
MIT is an acronym for the Massachusetts Institute of Technology. It is a prestigious private research university in Cambridge, Massachusetts that was founded in 1861. It is organized into five Schools: architecture and planning; engineering; humanities, arts, and social sciences; management; and science. MIT’s impact includes many scientific breakthroughs and technological advances. Their stated goal is to make a better world through education, research, and innovation.
Join us on Patreon! https://www.patreon.com/MichaelLustgartenPhD
Discount Links:
Oral Microbiome: https://www.bristlehealth.com/?ref=michaellustgarten.
Enter Code: ConquerAging.
At-Home Metabolomics: https://iollo.com?ref=michael-lustgarten.
NAD+ Quantification: https://www.jinfiniti.com/intracellular-nad-test/
Use Code: ConquerAging At Checkout.
Green Tea: https://www.ochaandco.com/?ref=conqueraging.
Epigenetic Testing: https://trudiagnostic.com/?irclickid=U-s3Ii2r7xyIU-LSYLyQdQ6…M0&irgwc=1

The biology underpinning a rare genetic mutation that allows its carrier to live virtually pain-free, heal more rapidly and experience reduced anxiety and fear, has been uncovered by new research from UCL.
The study, published in Brain, follows up the team’s discovery in 2019 of the FAAH-OUT gene and the rare mutations that cause Jo Cameron to feel virtually no pain and never feel anxious or afraid. The new research describes how the mutation in FAAH-OUT “turns down” FAAH gene expression, as well as the knock-on effects on other molecular pathways linked to wound healing and mood. It is hoped the findings will lead to new drug targets and open up new avenues of research in these areas.
Jo, who lives in Scotland, was first referred to pain geneticists at UCL in 2013, after her doctor noticed that she experienced no pain after major surgeries on her hip and hand. After six years of searching, they identified a new gene that they named FAAH-OUT, which contained a rare genetic mutation. In combination with another, more common mutation in FAAH, it was found to be the cause of Jo’s unique characteristics.