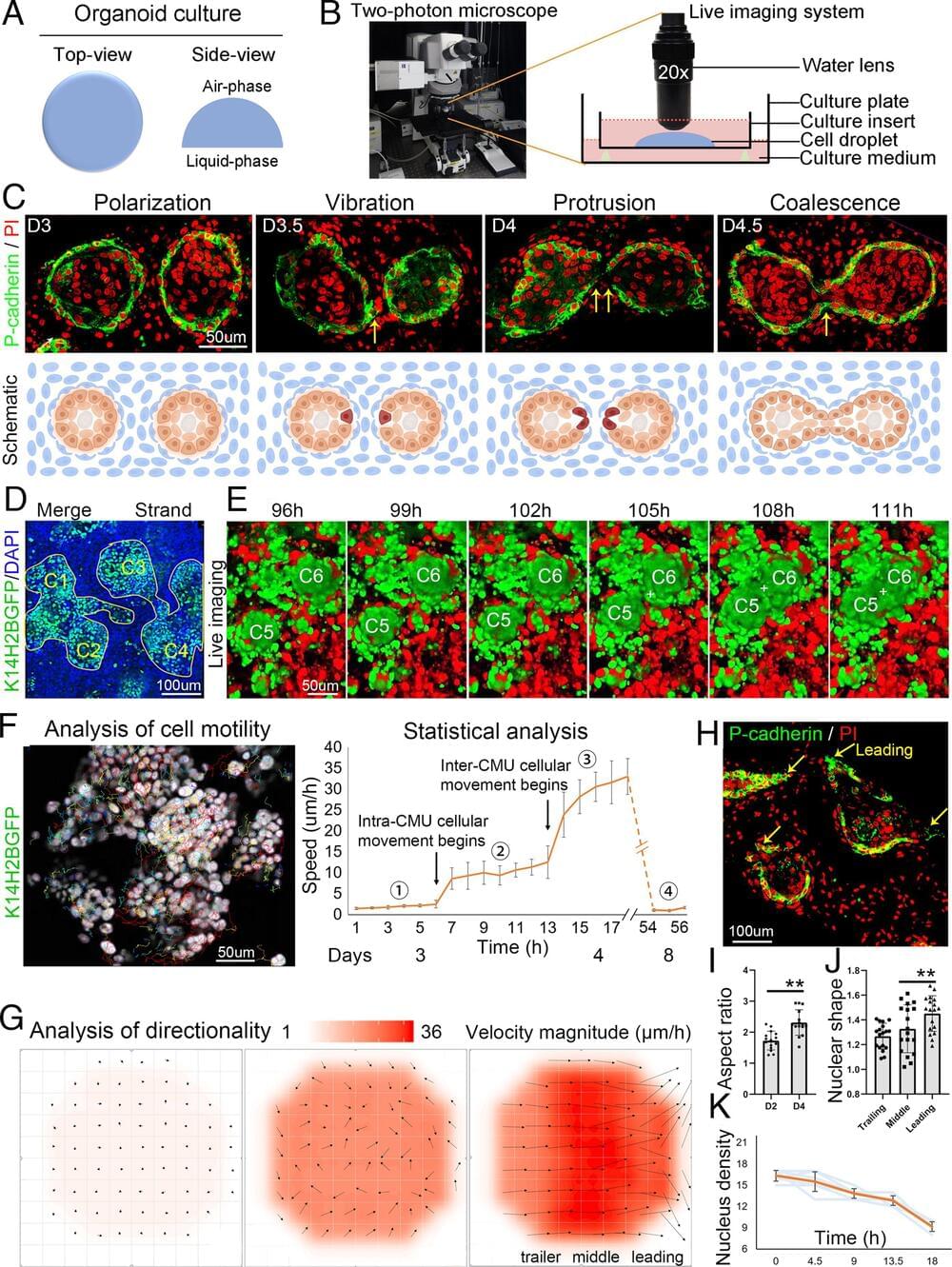
Stem cells in organoids self-organize into tissue patterns with unknown mechanisms. Here, we use skin organoids to analyze this process. Cell behavior videos show that the morphological transformation from multiple spheroidal units with morphogenesis competence (CMU) to planar skin is characterized by two abrupt cell motility–increasing events before calming down. The self-organizing processes are controlled by a morphogenetic module composed of molecular sensors, modulators, and executers. Increasing dermal stiffness provides the initial driving force (driver) which activates Yap1 (sensor) in epidermal cysts. Notch signaling (modulator 1) in epidermal cyst tunes the threshold of Yap1 activation. Activated Yap1 induces Wnts and MMPs (epidermal executers) in basal cells to facilitate cellular flows, allowing epidermal cells to protrude out from the CMU. Dermal cell–expressed Rock (dermal executer) generates a stiff force bridge between two CMU and accelerates tissue mixing via activating Laminin and β1-integrin. Thus, this self-organizing coalescence process is controlled by a mechano-chemical circuit. Beyond skin, self-organization in organoids may use similar mechano-chemical circuit structures.