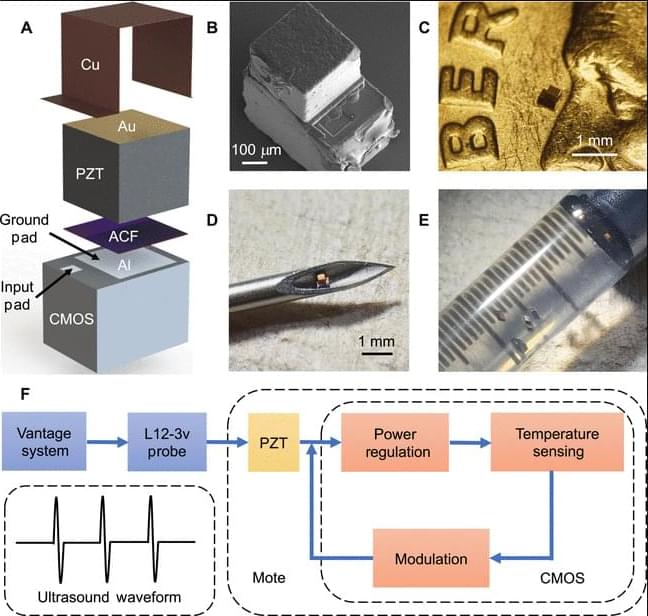
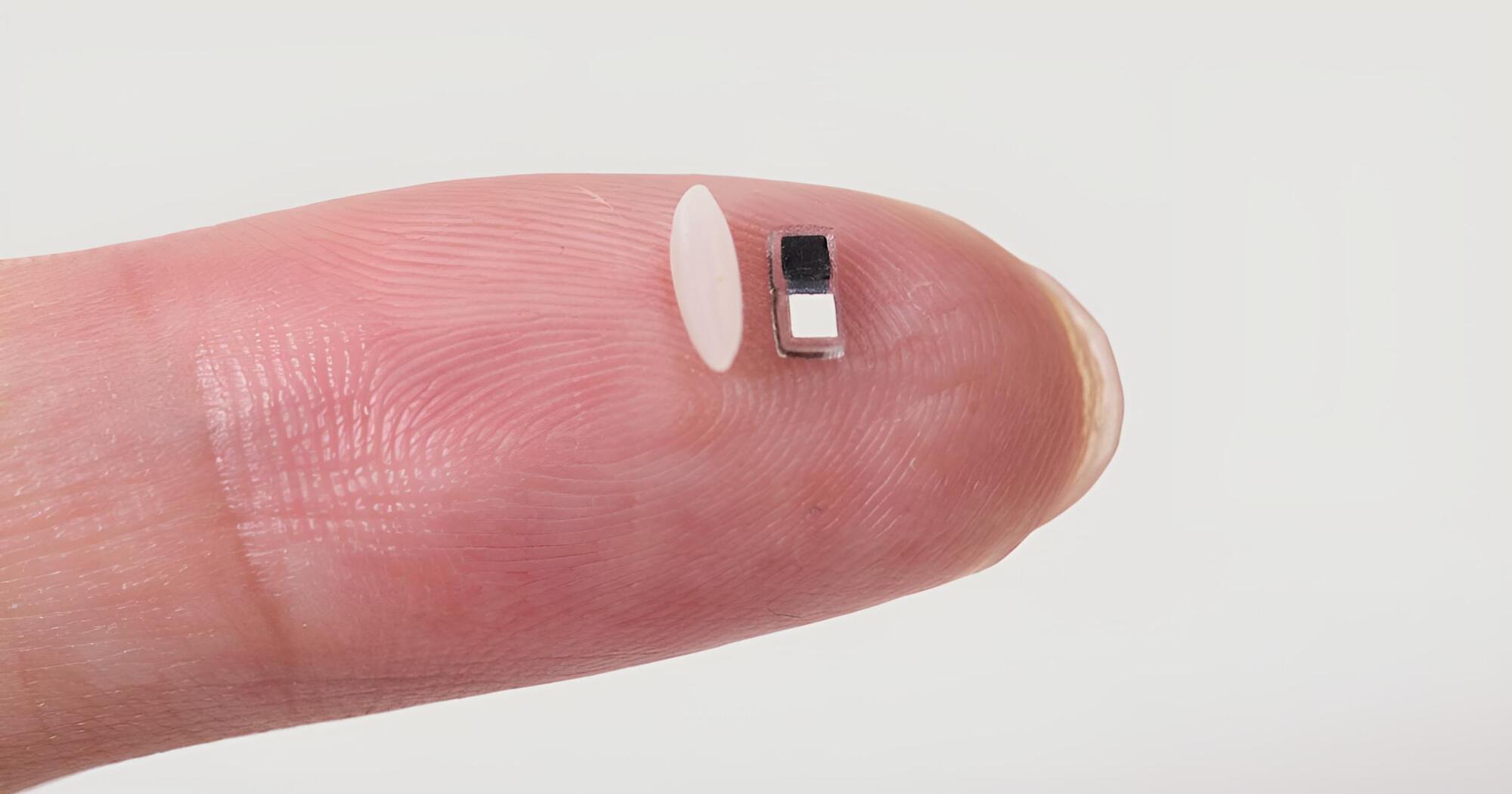
This paper presents a sub–0.1-mm3, fully integrated, injectable ultrasonic wireless device for biomedical temperature sensing.


The potential role of vitamin D in preventing and treating colorectal cancer (CRC) has attracted growing research interest – especially as CRC rates are rising, particularly among younger adults. This isn’t a new area of study. Low vitamin D levels have long been linked to a higher risk of developing colorectal cancer.
One large study involving over 12,000 participants found that people with low blood levels of vitamin D had a 31 per cent greater risk of developing CRC compared to those with higher levels. Similarly, another study reported a 25 per cent lower CRC risk among individuals with high dietary vitamin D intake.
Data from the Nurses’ Health Study – a long-term investigation of American nurses – showed that women with the highest vitamin D intake had a 58 per cent lower risk of developing colorectal cancer compared to those with the lowest intake.


The common cold sore virus, which is often caught in childhood, usually stays in the body for life—quietly dormant in the nerves. Now and then, things like stress, illness or injury can trigger it, bringing on a cold sore in some people. But this same virus—called herpes simplex virus type 1—may also play an important role in something far more serious: Alzheimer’s disease.
Over 30 years ago, my colleagues and I made a surprising discovery. We found that this cold sore virus can be present in the brains of older people. It was the first clear sign that a virus could be quietly living in the brain, which was long thought to be completely germ-free—protected by the so-called “blood-brain barrier.”
Then we discovered something even more striking. People who have a certain version of a gene (called APOE-e4) that increases their risk of Alzheimer’s, and who have been infected with this virus, have a risk that is many times greater.
A new gene therapy reversed heart failure in pigs by repairing heart function through cBIN1, showing major promise for future treatment.
A new gene therapy has been shown to reverse the effects of heart failure and restore heart function in a large animal model. The treatment increases the heart’s ability to pump blood and significantly improves survival rates. A paper describing the results calls it “an unprecedented recovery of cardiac function.”
Heart failure is currently irreversible. Without a heart transplant, most treatments aim only to reduce the heart’s workload and slow the progression of the disease. If this gene therapy produces similar outcomes in future clinical trials, it could offer a way to repair the hearts of one in four people expected to develop heart failure during their lifetime.

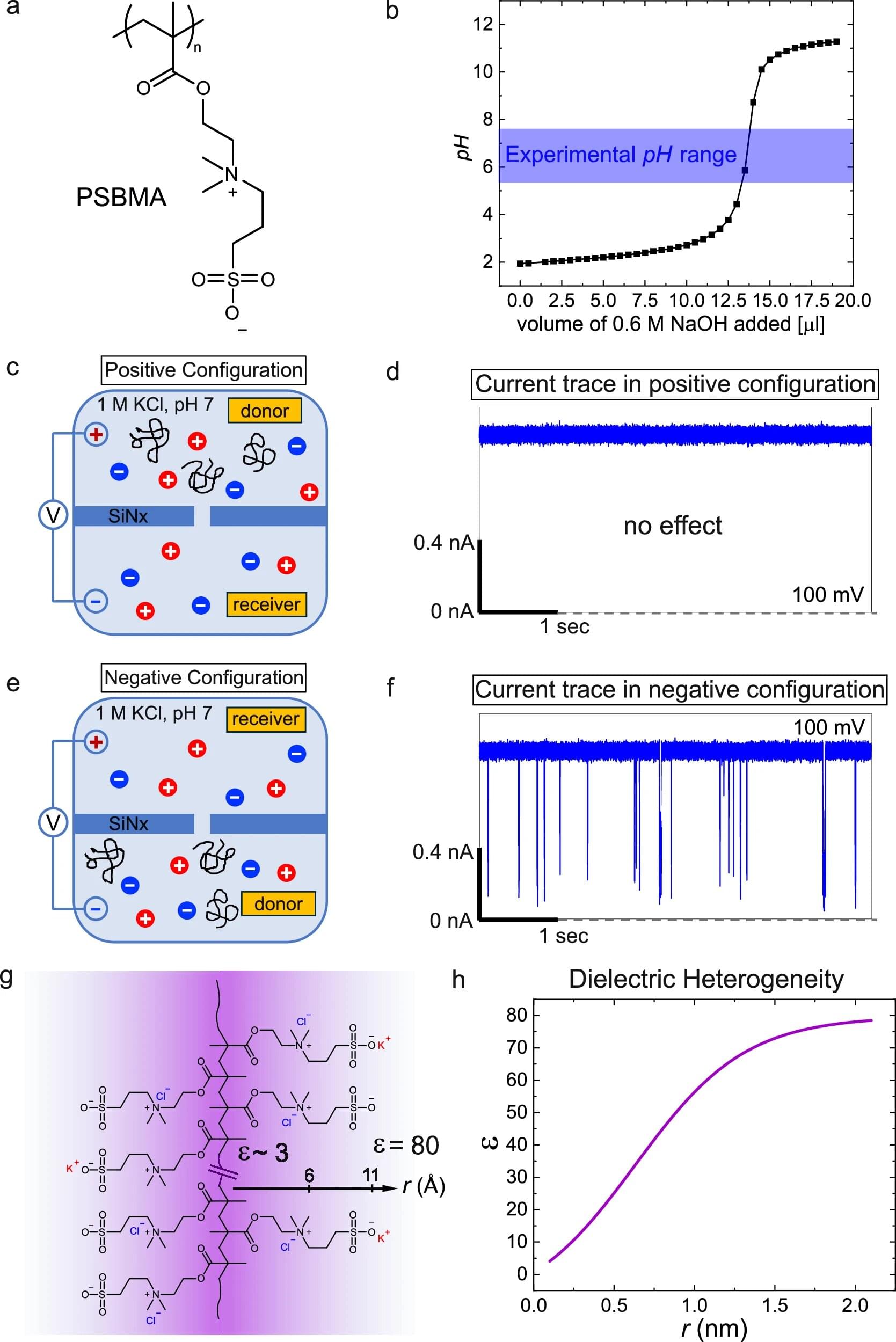
A new study led by a pair of researchers at the University of Massachusetts Amherst turns long-held conventional wisdom about a certain type of polymer on its head, greatly expanding understanding of how some of biochemistry’s fundamental forces work. The study, released recently in Nature Communications, opens the door for new biomedical research running the gamut from analyzing and identifying proteins and carbohydrates to drug delivery.
The work involves a kind of polymer made up of neutral polyzwitterions. Because they have a neutral electrical charge, polyzwitterions are not expected to respond to an electric field. However, the team found not only that certain neutral polyzwitterions behave as if they were charged, but also that the electric field surrounding polyzwitterions, once thought to be uniform, varies in strength.
“My interest is in the proteins and amino acids, which are the building blocks for protein, inside our body’s cells,” says Yeseul Lee, lead author and graduate student in polymer science and engineering at UMass Amherst.
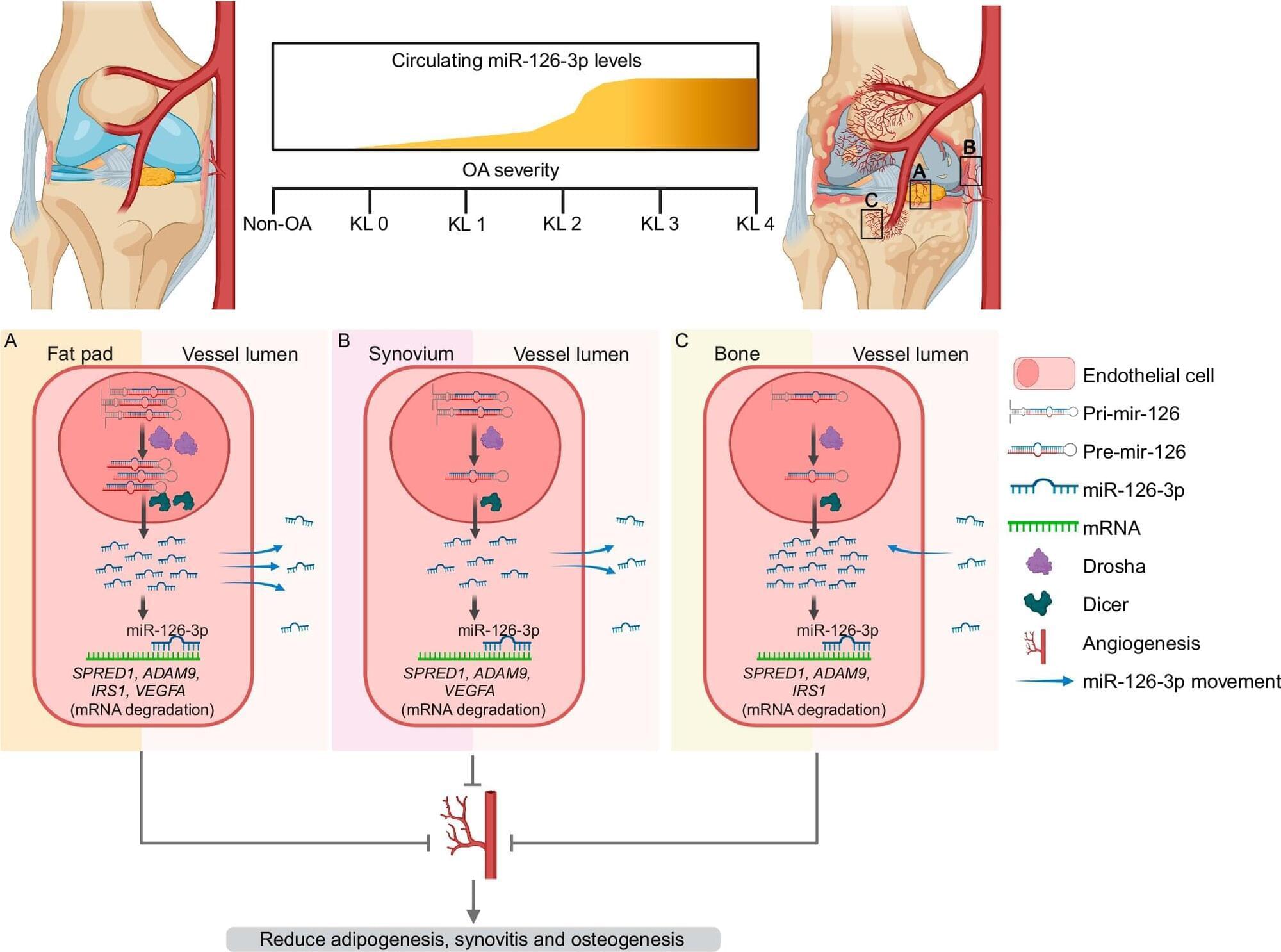
The number of people suffering from osteoarthritis is expected to top 1 billion by 2050. The biggest risk factor for the prevalent, often painful, chronic joint disease is aging. And like aging, there is currently no way to stop it.
A discovery by scientists at Henry Ford Health + Michigan State University Health Sciences could pave the way for new breakthroughs in detecting and treating the disease. Their findings were recently published in Nature Communications.
“Our hope is that this discovery will one day allow doctors to catch the disease earlier and intervene before significant joint damage occurs,” said Shabana Amanda Ali, Ph.D., a Henry Ford Health assistant scientist and senior author of the paper. “Osteoarthritis is so complex and so heterogeneous that even with decades of research there hasn’t been a single therapeutic.”