Patients regained movement in the experimental phase of polylaminin, capable of regenerating the spinal cord in people who suffered organ rupture


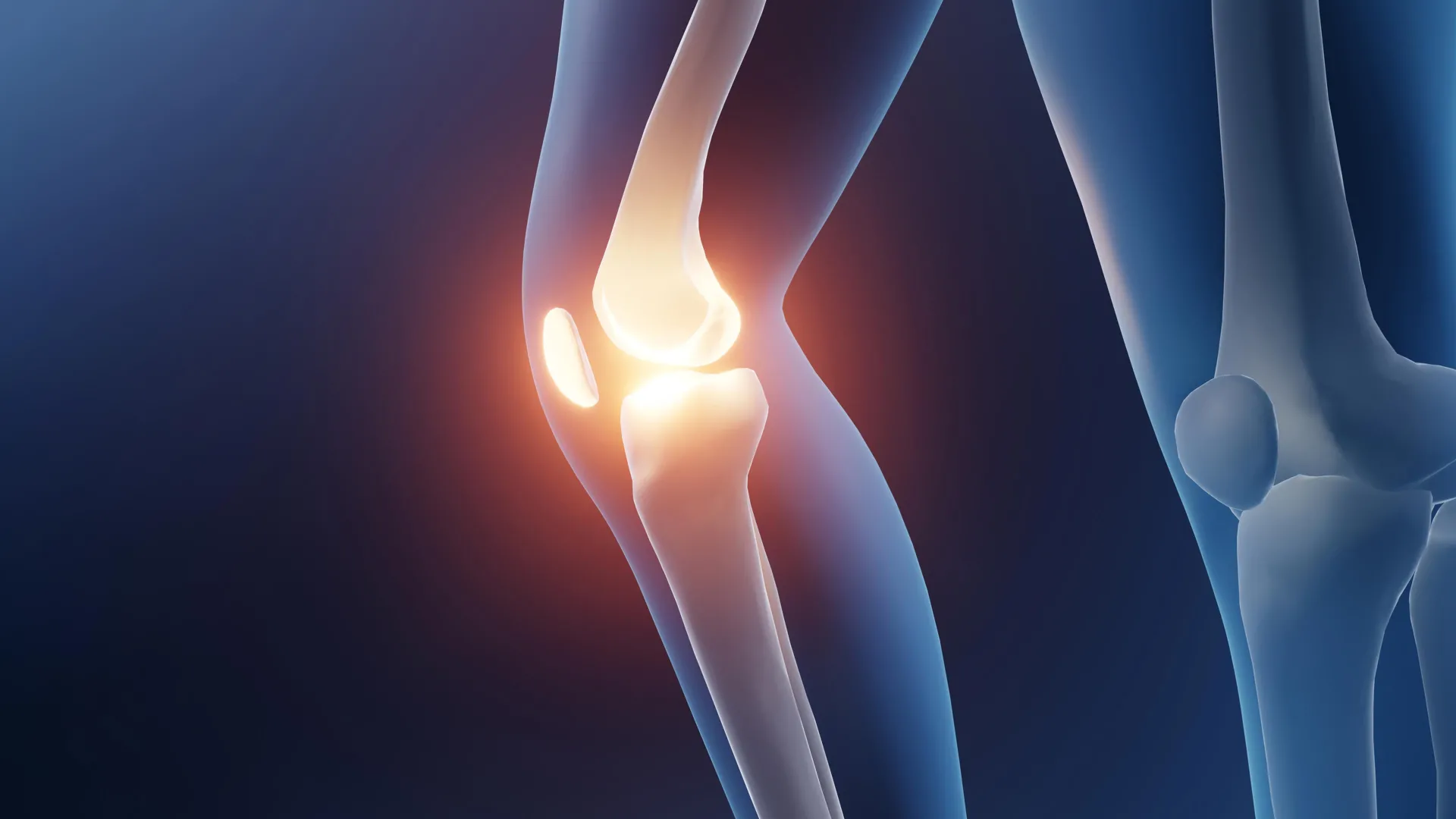
Cambridge scientists have created a breakthrough material that can sense tiny chemical changes in the body, such as the increased acidity during an arthritis flare-up, and release drugs exactly when and where they’re needed. By mimicking cartilage while delivering medication, this smart gel could ease pain, reduce side effects, and provide continuous treatment for millions of arthritis sufferers.
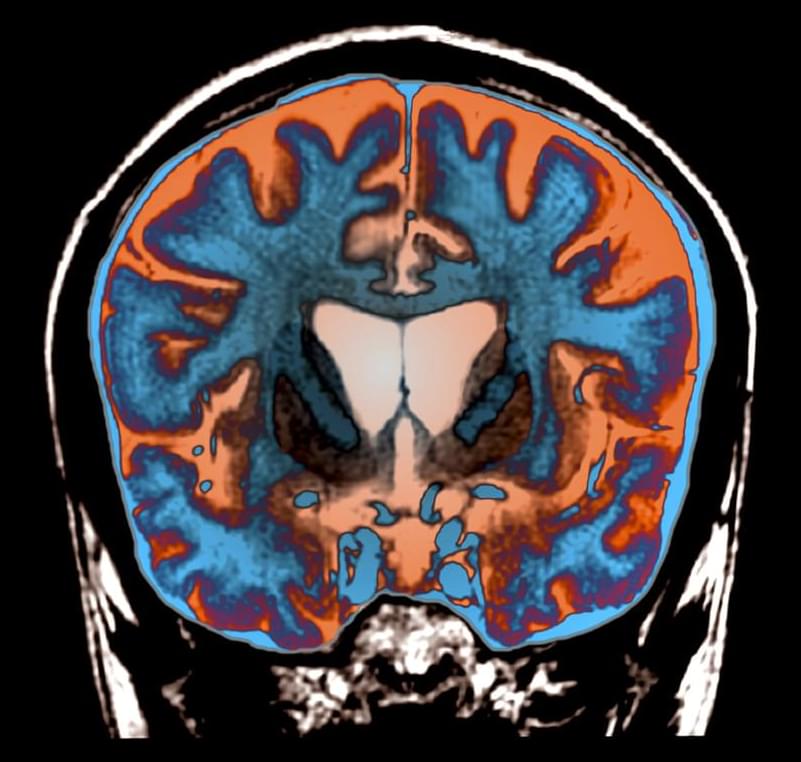
Even hearing the phrase “Huntington’s disease” will make a room suddenly somber. So the joy that accompanied a recent announcement of results of an experimental gene therapy for the deadly diseases signaled an unfamiliar sense of hope.
In a small clinical trial, brain injections of a virus that codes for a tiny segment of RNA may have prevented the formation of the rogue proteins that make Huntington’s so devastating. The early results, announced September 24 in a news release, show that over three years, the treatment slowed Huntington’s progression by up to 75 percent. While not a cure, the treatment could potentially give people living with Huntington’s disease, who might otherwise face early disability and death, the gift of many more years of life.
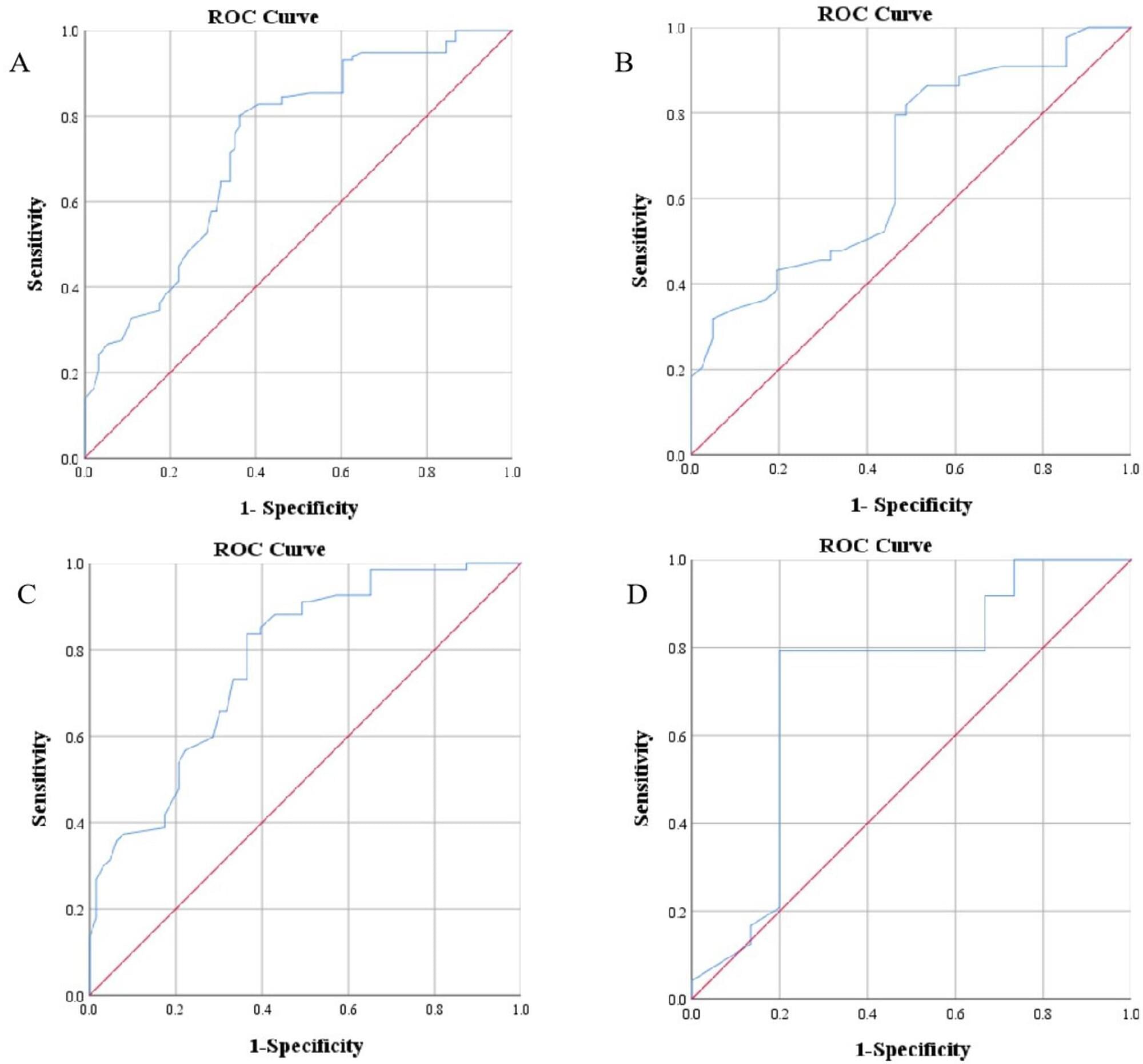
Early oral propranolol for infantile hemangioma (IH) can improve the success rate. However, few studies have been conducted on the specific age threshold for initiating treatment with propranolol. This study aimed to determine the cutoff value of treatment success for IH with oral propranolol.
This was a retrospective study involving 207 patients with IH and clinical data. The primary outcome measure was treatment success with oral propranolol at months 6 to 12.
Multivariate analysis showed that age at treatment initiation (P 0.001), high-risk IH (P = 0.002), and segmental IH (P = 0.012) were independent risk factors for treatment unsuccess with oral propranolol at months 6 to 12. Receiver operating characteristic (ROC) curve analysis indicated that the cutoff value for age at treatment initiation was 69.5 days for all patients, 65.5 days for patients with segmental IH or high-risk IH, and 93.5 days for patients with non-high-risk and nonsegmental IH. Patients who started treatment before 69.5 days had a success rate of 73.8%, which was higher than the 28.4% success rate of patients who started treatment after 69.5 days. The median time to treatment success for patients who started treatment before 69.5 days was 11.5 months, which was shorter than the 15 months for patients who started treatment after 69.5 days.
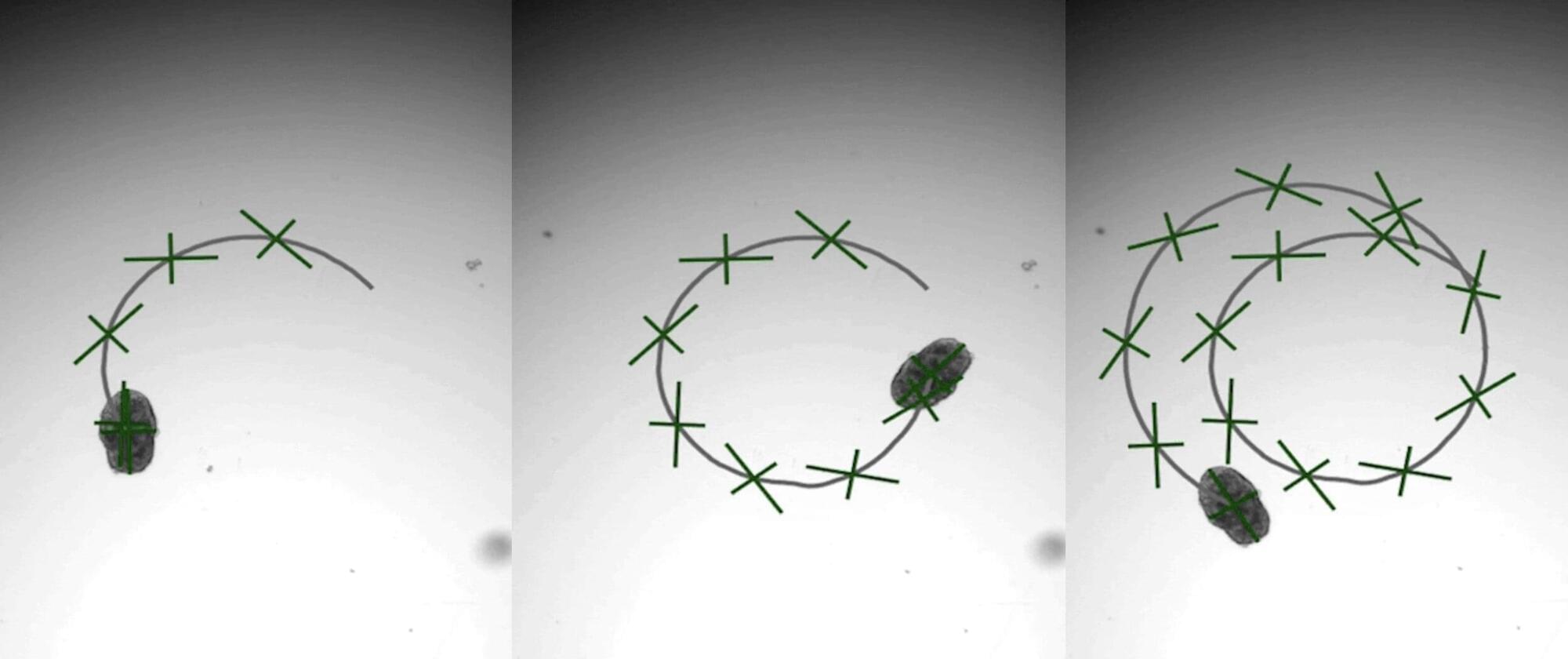

A brand-new engineering approach to generate “designer” biological robots using human lung cells is underway in Carnegie Mellon University’s Ren lab. Referred to as AggreBots, these microscale living robots may one day be able to traverse through the body’s complex environments to deliver desired therapeutic or mechanical interventions, once greater control is achieved over their motility patterns. In new research published in Science Advances, the group provides a novel tissue engineering platform capable of achieving customizable motility in AggreBots by actively controlling their structural parameters.
Biobots are microscopic, man-made biological machines capable of autonomous movement and programmability to perform specific tasks or behaviors. Previously, enabling biobots’ motility has been centered around using muscle fibers, which allow them to move by contracting and relaxing like real muscles.
A novel, alternative mechanism of actuation can be found by using cilia, the nanoscopic, hair-like, organic propellers that continuously move fluids in the body (like in the lungs) and help some aquatic creatures, like Paramecium or comb jellies, swim. However, a reliable way to control the exact shape and structure of a cilia-powered biobot (CiliaBot, for short), and thereby its motility outcome, has proven difficult to come by.
Shaping The Future Of Personalized Medicine — Professor Dragan[ ](https://www.facebook.com/PrimoracDragan?__cft__[0]=AZWpslTHjsy1a1kjedsti2RJw9yv6FhOXDFg2kyiufa2-D4Gk8TYoTy6HPaDPGARaq1EESF8mpBiV9Jjt2gpkh8Np3gpvzqTNu4cOTW-m31Hn4MVmEFyC6gnP5_-bMEdn1Gn81MUYh3llD5MqtPqF8dPWOZxq1Oo7MbC2g5664Of2FI4tc98YxJrFewUmig_tH0&__tn__=-]K-R)Primorac MD, PhD — Founder, St. Catherine Specialty Hospital
Professor Dragan Primorac, MD., PhD (https://www.draganprimorac.com/) is a globally recognized physician-scientist whose work spans personalized medicine, regenerative therapies, and forensic genetics.
From 2003 to 2009 Prof. Primorac served as the Minister of Science, Education and Sports of the Republic of Croatia. The Ministry of Science and Education of Croatia is the ministry in the Government of Croatia which is in charge of primary, secondary and tertiary education, research institutions and sports (https://mzom.gov.hr/en).
Prof. Primorac is the Founder of St. Catherine Specialty Hospital in Zagreb Croatia (https://www.stcatherine.com/), the official hospital of the Croatian Olympic Committee as well as the official hospital of the Croatian Football Federation. St. Catherine Hospital is affiliated with four medical schools and the Ministry of Science and Education recently announced that the St. Catherine Hospital became Scientific Center of Excellence for the Personalized Medicine\.

Harnessing the power of AI, a research team at the MRC-University of Glasgow Center for Virus Research has launched Viro3D —the most comprehensive database of human and animal virus protein structure predictions in the world.
The free and searchable AI-powered database offers a completely new, in-depth perspective on viruses, allowing us to quickly learn more about their origins and evolution.
Although virus particles are the most abundant biological entities on our planet, these tiny structures remain among the least well-understood. Insights into the key protein structures within viruses have, until now, only been achieved through slow and laborious research work, a pace that has impacted our ability to develop treatments and vaccines at speed.
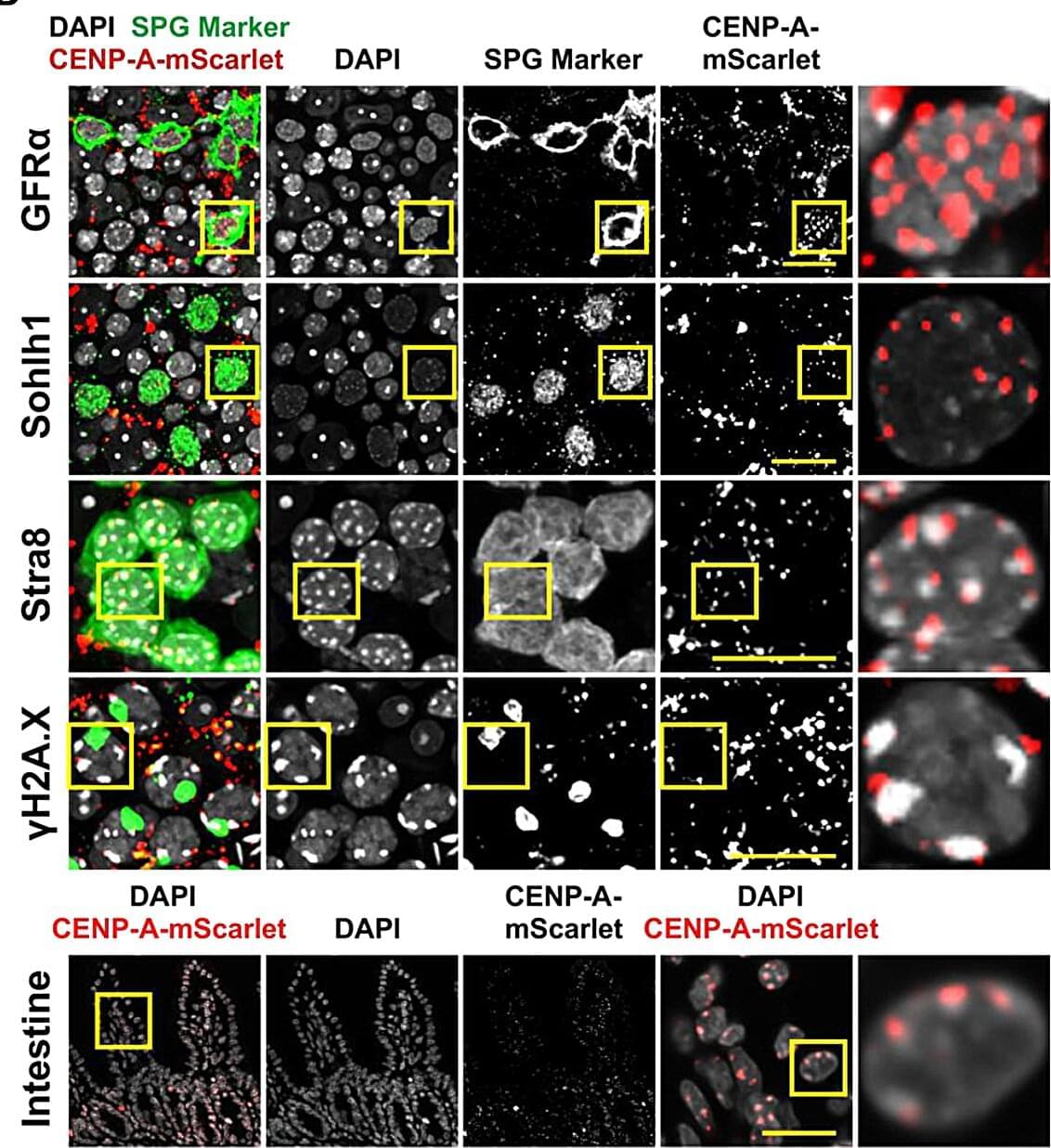
When a sperm meets an egg, a lot has to go right for an embryo to develop into a complete organism. One critical step of early development is the reorganization of parental DNA to form a new unified genome, before the embryo can undergo its first cell division.
Scientists have long known that sperm and eggs bundle their DNA differently. But it’s been assumed that their centromeres—the special regions of each chromosome that act like handles to pull DNA apart during cell division —were essentially the same. That assumption rested on the presence of centromere protein A, or CENPA, a unique histone protein that marks centromeres and preserves their identity across each cell division and across generations.
Because CENPA acts like a molecular tag, preserving these sites as “do not erase” regions of the genome, the centromeres were thought to be functionally indistinguishable between maternal and paternal chromosomes.

The International Advisory Committee on Clinical Trials led a multinational panel updating the McDonald criteria, adding the optic nerve as a fifth anatomical location and allowing specific magnetic resonance imaging (MRI) and cerebrospinal fluid (CSF) markers to support diagnosis without mandatory dissemination in time in defined scenarios.
Multiple sclerosis diagnosis has long required proof that lesions occur in different places and at different times, with MRI and CSF biomarkers gradually shortening time to treatment.
Previous revisions improved sensitivity and specificity across ages and regions, yet misdiagnosis risk still persists, especially with overlapping conditions and when access to specialized tests is limited.

In a world of information overload, it can feel soothing to stick your head in the sand.
Don’t want to hear what the doctor might say? It’s easy not to make a follow-up appointment. Did a favorite political candidate say something you disagreed with? The evidence can disappear with a flick of a finger.
According to psychologists, avoiding information when it’s uncomfortable is a common adult behavior, often referred to as the “Ostrich Effect.”