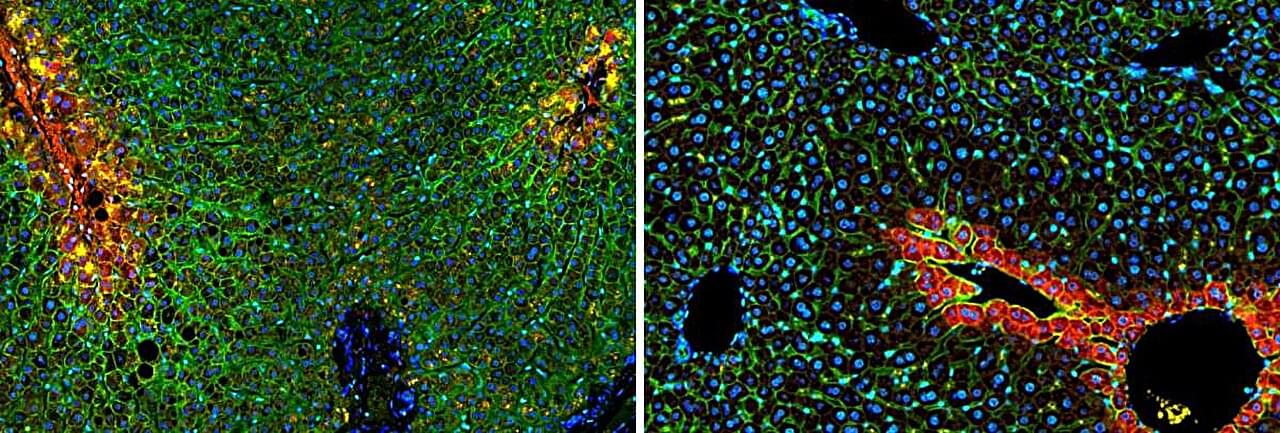
If scientists could shrink themselves to microscopic size and take a journey through the human body—like the submarine crew in the 1966 science fiction classic “Fantastic Voyage”—one of their first stops would no doubt be the liver. The unique structure of our largest internal organ comprises small, hexagonal functional units called lobules, each carrying out more than 500 functions simultaneously. Studies from the 1970s and 1980s revealed that liver cells divide these many tasks among themselves according to their location within each subunit; however, the technology available at the time provided only a blurred picture of this division of labor.
In a new study published in Nature, scientists from the Weizmann Institute of Science, together with colleagues at Sheba Medical Center and the Mayo Clinic, present the first genetic atlas of a healthy human liver at a resolution of 2 microns. The findings show that the division of labor in the human liver differs from that of other mammals and is more extensive than previously recognized, helping explain why certain regions of the liver are particularly vulnerable to fatty liver disease.
In recent years, technological advances have made it possible to identify which genes are active in each individual cell while also mapping the cells’ precise spatial positions within the tissue. Still, a comprehensive map of functional division in the human liver remained elusive, largely due to the difficulty of obtaining tissue samples from healthy donors.