A new University of California San Diego School of Medicine study offers a unified biological model to explain how genetic predispositions and environmental exposures converge to cause autism spectrum disorder (ASD).
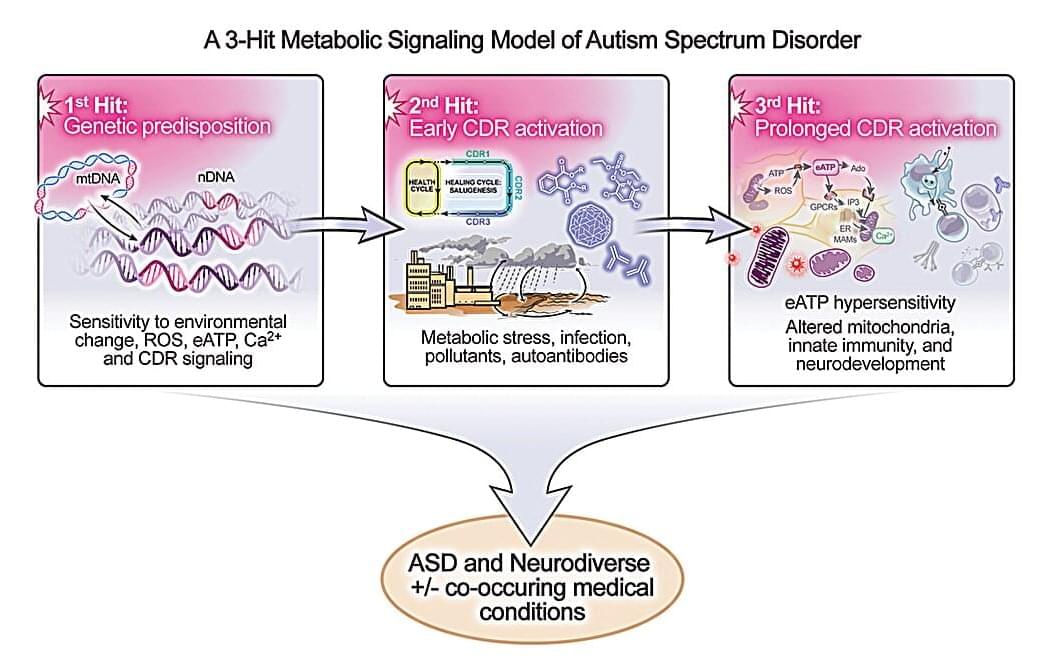
The study, published in Mitochondrion, describes a “three-hit” metabolic signaling model that reframes autism as a treatable disorder of cellular communication and energy metabolism. The model also suggests that as many as half of all autism cases might be prevented or reduced with prenatal and early-life interventions.
“Our findings suggest that autism is not the inevitable result of any one gene or exposure, but the outcome of a series of biological interactions, many of which can be modified,” said study author Robert K. Naviaux, M.D., Ph.D., professor of medicine, pediatrics and pathology at UC San Diego School of Medicine.