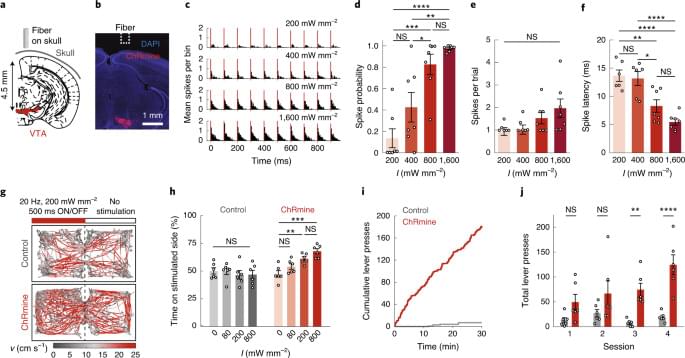
By combining combine genetic and electrical engineering, scientists have developed a new technique for wiring electronics into living matter.
Category: genetics – Page 296

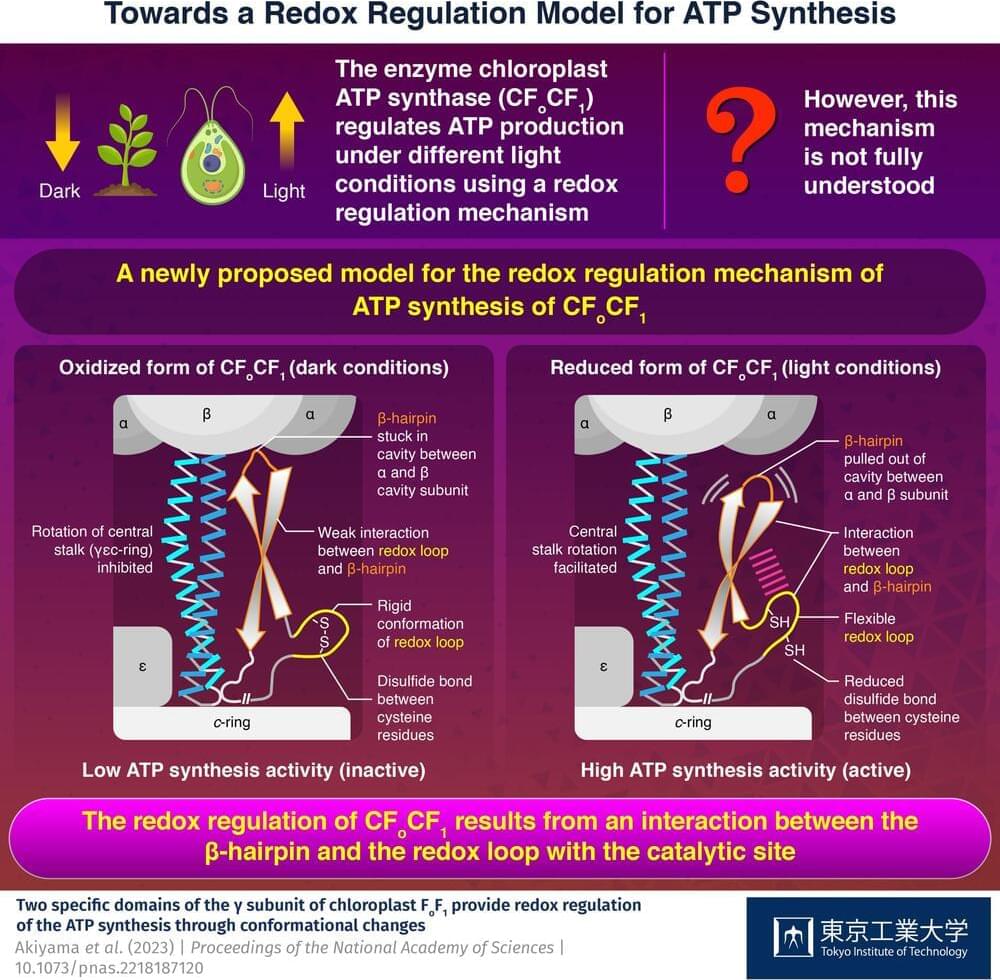
Researchers uncover how photosynthetic organisms regulate and synthesize ATP
ATP, the compound essential for the functioning of photosynthetic organisms such as plants, algae, and cyanobacteria, is produced by an enzyme called “chloroplast ATP synthase” (CFoCF1). To control ATP production under varying light conditions, the enzyme uses a redox regulatory mechanism that modifies the ATP synthesis activity in response to changes in the redox state of cysteine (Cys) residues, which exist as dithiols under reducing (light) conditions, but forms a disulfide bond under oxidizing (dark) conditions. However, this mechanism has not yet been fully understood.
Now, in a study published in the Proceedings of the National Academy of Sciences, a team of researchers from Japan led by Prof. Toru Hisabori from Tokyo Institute of Technology (Tokyo Tech) has uncovered the role of the amino acid sequences present in CFoCF1, revealing how the enzyme regulates ATP production in photosynthetic organisms.
To understand how the conformation of the amino acids present in CFoCF1 contributes to the redox regulation mechanism, the researchers used the unicellular green alga, Chlamydomonas reinhardtii, to produce the enzyme. “By leveraging the powerful genetics of Chlamydomonas reinhardtii as a model organism for photosynthesis, we conducted a comprehensive biochemical analysis of the CFoCF1 molecule,” explains Prof. Hisabori.
Researchers provide a framework for unifying and categorizing neurodegenerative diseases based on eight hallmarks
In a recent study published in Cell, researchers presented eight hallmarks of neurodegenerative diseases (NDDs), their in vivo biomarkers, and interactions to help categorize NDDs and specify patients within a specific NDD.
Despite being linked to rare genetic forms, all eight NDD hallmarks (cellular/molecular processes) also contribute to sporadic NDDs. In addition, they contribute to neuronal loss in preclinical (animal) models and NDD patients, manifesting as an altered molecular (hallmark) biomarker.
An NDD patient could have defects in multiple NDD hallmarks. However, the primary NDD hallmark depends on the NDD insult and the neuronal susceptibility and resilience, i.e., one’s ability to handle insults in the affected brain region.
Cellular senescence plays a significant role in cerebral tumors
Glioblastomas are the most common malignant tumors of the adult brain. They resist conventional treatment, including surgery, followed by radiation therapy and chemotherapy. Despite this armamentarium, glioblastomas inexorably recur.
In a new study published in Nature Communications, Isabelle Le Roux (CNRS) and her colleagues from the “Genetics and development of brain tumors” team at Paris Brain Institute have shown that the elimination of senescent cells, i.e., cells that have stopped dividing, can modify the tumor ecosystem and slow its progression. These results open up new avenues for treatment.
Glioblastoma, the most common adult brain cancer, affects 2 to 5 in 100,000 individuals. While the incidence of the disease is highest in those between 55 and 85 years old, it is increasing in all age groups. This effect can’t be attributed to improved diagnostic techniques alone, suggesting the influence of environmental factors hitherto unidentified.
Dr. Abdelali Haoudi, PhD — KAIMRC — Advancing Biomedical R&D & Clinical Development In Saudi Arabia
Advancing Biomedical R&D & Clinical Development In Saudi Arabia — Dr. Abdelali Haoudi, Ph.D., Managing Director, Biotechnology Park, King Abdullah International Medical Research Center, Ministry of National Guard Health Affairs.
Dr. Abdelali Haoudi, Ph.D. (https://kaimrc-biotech.org.sa/dr-abdelali-haoudi/) currently leads Strategy and Business Development functions, and is also Managing Director of the Biotechnology Park, at King Abdullah International Medical Research Center, at the Ministry of National Guard Health Affairs. He is also Distinguished Scholar at Harvard University-Boston Children’s Hospital.
Dr. Haoudi is an international Research & Development and Innovation Executive with over 25 years experience, having held several senior positions in Research and Development and Innovation. He has vast experience in science and technology policy development, strategy and business development, corporate development and international partnerships development.
Dr. Haoudi has held several senior and prestigious positions in the academia, government and private sectors globally, including North America, North Africa, Europe and Middle East. Some of the key positions include Founding Vice President for Research, Executive Director for Biomedical Research Institute, and Chairman, at the National Research Fund at Qatar Foundation.
Dr. Haoudi was also Research Professor of cancer and infectious diseases at the University of Virginia and Eastern Virginia Medical School and a fellow of the US National Institutes of Health (NIH). He has held several other positions in elite research and education institutions including Visiting Professor at Harvard Medical School (USA) and a fellow at Institute Pasteur (France).
Epigenetic and social factors both predict aging and health, but new research suggests one might be stronger
Can we objectively tell how fast we are aging? With a good measure, scientists might be able to change our rate of aging to live longer and healthier lives. Researchers know that some people age faster than others and have been trying to concisely measure the internal physiological changes that lead to deteriorating health with age.
For years, researchers have been using clinical factors normally collected at physicals, like hypertension, cholesterol and weight, as indicators to predict aging. The idea was that these measures could determine whether someone is a fast or slow ager at any point in their life cycle. But more recently, researchers have theorized that there are other biological markers that reflect aging at the molecular and cellular level. This includes modifications to a person’s genetic material itself, or epigenetics.
While each person has a genetic makeup that largely does not change over their lifetime, chemical changes to their genetic material that occur throughout life can change which genes are turned on or off and lead to more rapid aging. These changes typically involve the addition of methyl groups to DNA and are influenced by social and environmental exposures, such as adverse childhood experiences, smoking, pollution and depression.
Telomeres Found To Encode Two Proteins, Potentially Transforming Cancer Research
Telomeres – the protective caps at the tips of chromosomes – can encode two proteins, something that was previously thought impossible, new research has suggested. The discovery of genetic information coding for these proteins, one of which is elevated in some human cancers, could have huge ramifications for the fields of health, medicine, and cell biology.
“Discovering that telomeres encode two novel signaling proteins will change our understanding of cancer, aging, and how cells communicate with other cells,” study author Jack Griffith, the Kenan Distinguished Professor of Microbiology and Immunology at the University of North Carolina at Chapel Hill, said in a statement.
“Based on our research, we think simple blood tests for these proteins could provide a valuable screen for certain cancers and other human diseases,” Griffith, who is also a member of the UNC Lineberger Comprehensive Cancer Center, added. “These tests also could provide a measure of ‘telomere health,’ because we know telomeres shorten with age.”
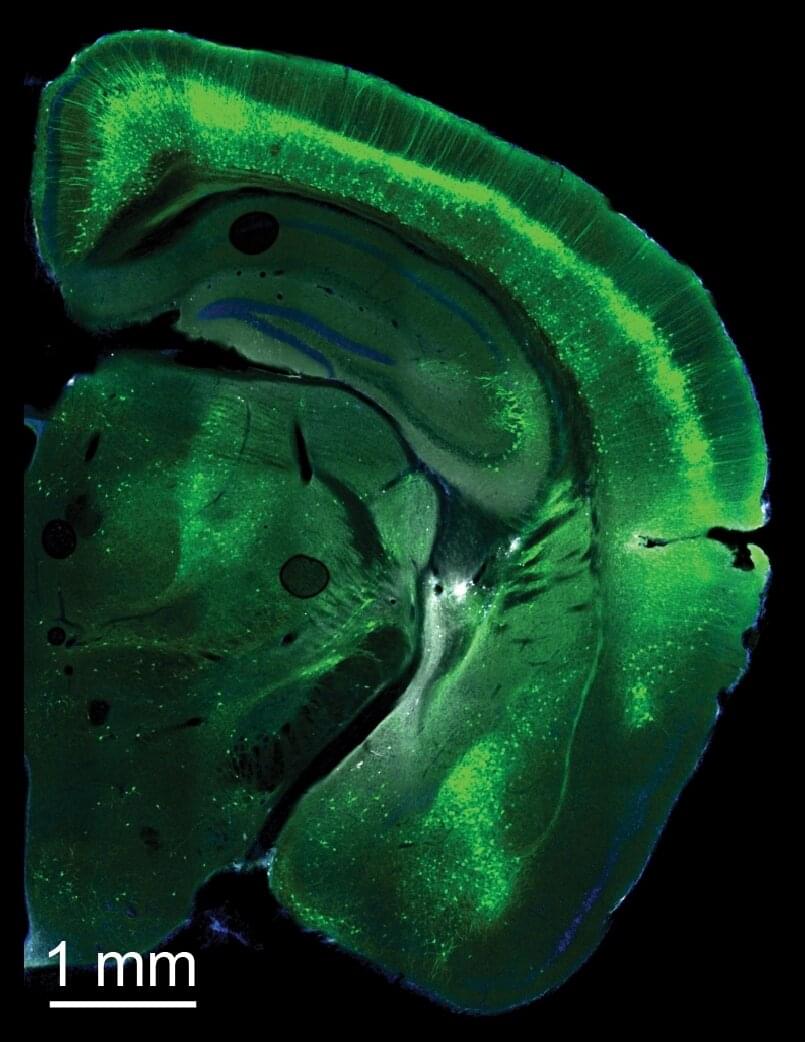
Study shows that distinct pyramidal cell types drive different patterns of cortical activity during decision-making
The outer layer of the brain, known as the cortex, is made of different types of neurons. Neuroscience studies suggest that these different neuron types have distinct functions, yet for a long time this was difficult to ascertain, due to the inability to examine and manipulate them in the brains of living beings.
In recent years, genetic techniques opened new possibilities for studying cells and their functions. Using some of these techniques, researchers at Forschungszentrum Jülich, RWTH Aachen University, Cold Spring Harbor Laboratory and other institutes in the United States closely examined the functions of different pyramidal cells, which are commonly found in the human cortex.
Their findings, published in Nature Neuroscience, suggest that distinct types of pyramidal cells drive patterns of cortical activity associated with different brain functions. The team’s study builds on some of their previous works focusing on neuronal activity in the cortex.