New data link low-dose mTOR inhibition to DNA protection in aging immune cells, reframing how rapamycin’s healthspan effects might work.


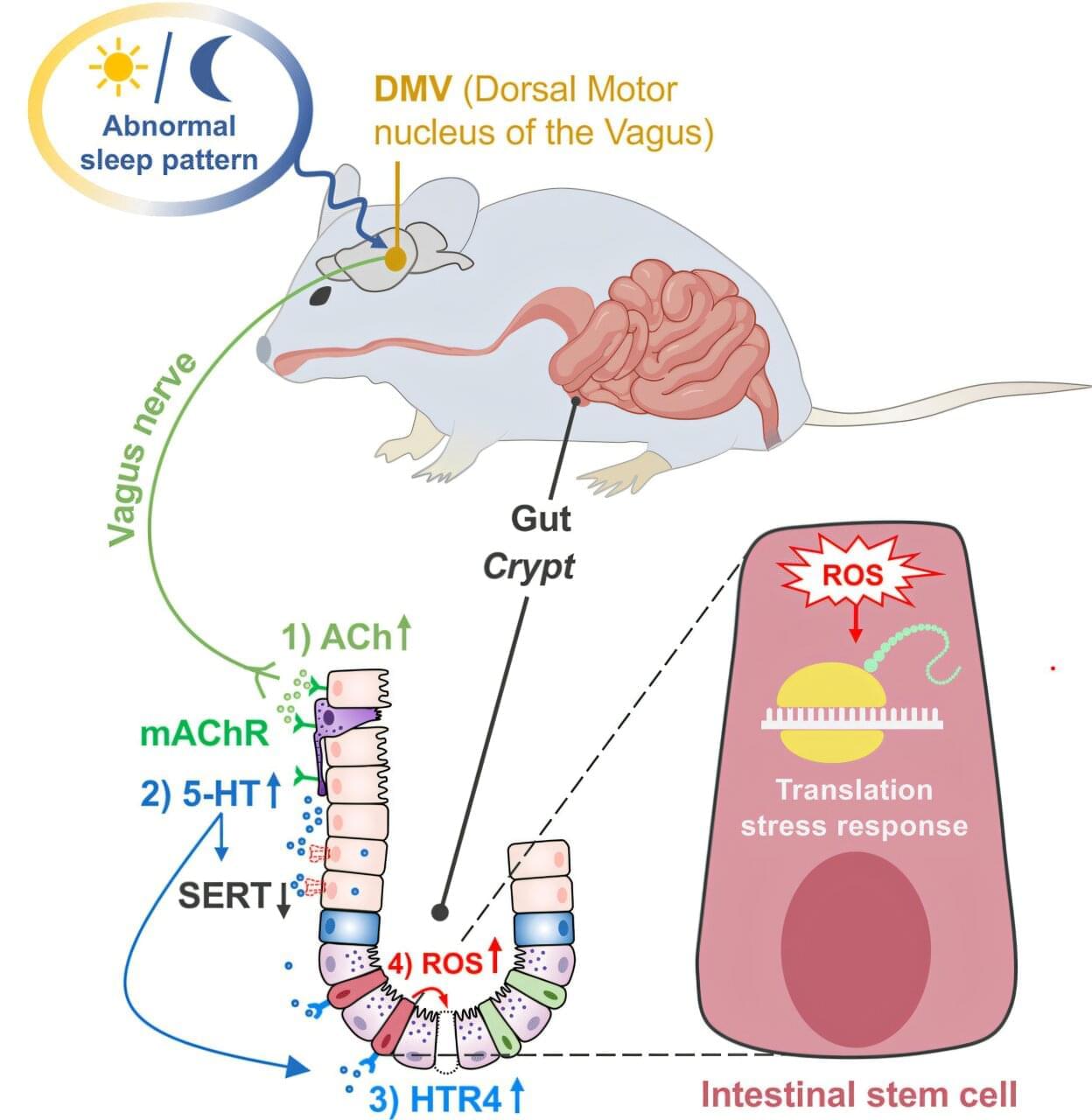
Chronic sleep disruption doesn’t just leave people tired and irritable. It may quietly undermine the gut’s ability to repair itself, increasing vulnerability to serious digestive diseases. A new study from the University of California, Irvine, the University of Chinese Academy of Sciences and the China Agricultural University reveals, step by step, how disturbed sleep causes the brain to send harmful signals to the intestines, ultimately damaging the stem cells responsible for maintaining a healthy gut lining.
The research uncovers a previously unknown biological chain reaction linking the brain’s sleep center to intestinal health. The findings are published in Cell Stem Cell and offer new insight into why people with chronic sleep problems are more likely to develop gastrointestinal disorders such as inflammatory bowel disease, diabetes-related gut complications and chronic inflammation.
Physicians have long known that irregular or insufficient sleep is associated with a wide range of health problems, from mood disorders to high blood pressure. Yet how changes in sleep can directly harm organs that do not sleep themselves, such as the intestines, has remained largely elusive. This study answers that question by tracing the damage from its neurological origins all the way to the gut’s regenerative machinery.

All life on Earth shares a common ancestor that lived roughly four billion years ago. This so-called “last universal common ancestor” (LUCA) represents the most ancient organism that researchers can study. Previous research on the last universal common ancestor has found that all the characteristics we see in organisms today, like having a cell membrane and a DNA genome, were already present by the time of this ancestor. So, if we want to understand how these foundational characteristics of life first emerged, then we need to be able to study evolutionary history prior to the last universal common ancestor.
In an article published in the journal Cell Genomics, scientists Aaron Goldman (Oberlin College), Greg Fournier (MIT), and Betül Kaçar (University of Wisconsin‑Madison) describe a method to do just that.
“While the last universal common ancestor is the most ancient organism we can study with evolutionary methods,” said Goldman, “some of the genes in its genome were much older.” The authors describe a type of gene family known as a “universal paralog,” which provides evidence of evolutionary events that occurred before the last universal common ancestor.

The trajectory of a storm, the evolution of stock prices, the spread of disease — mathematicians can describe any phenomenon that changes in time or space using what are known as partial differential equations. But there’s a problem: These “PDEs” are often so complicated that it’s impossible to solve them directly.
Mathematicians instead rely on a clever workaround. They might not know how to compute the exact solution to a given equation, but they can try to show that this solution must be “regular,” or well-behaved in a certain sense — that its values won’t suddenly jump in a physically impossible way, for instance. If a solution is regular, mathematicians can use a variety of tools to approximate it, gaining a better understanding of the phenomenon they want to study.
But many of the PDEs that describe realistic situations have remained out of reach. Mathematicians haven’t been able to show that their solutions are regular. In particular, some of these out-of-reach equations belong to a special class of PDEs that researchers spent a century developing a theory of — a theory that no one could get to work for this one subclass. They’d hit a wall.
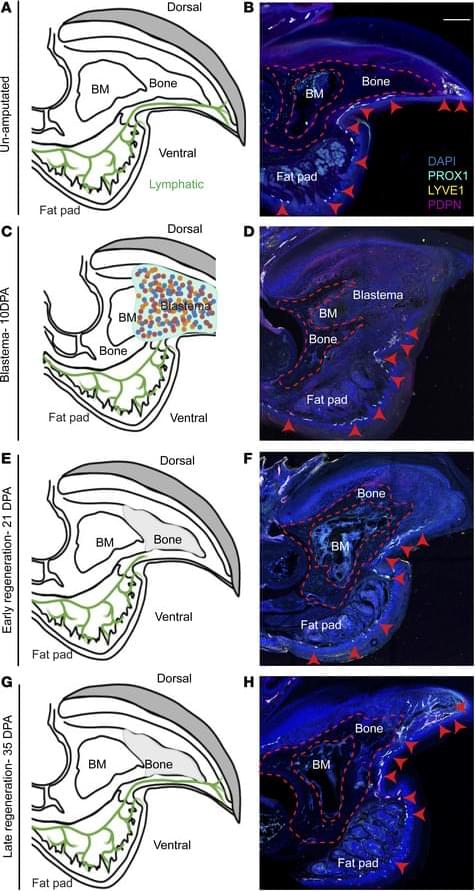
Matthijs Luxen et al. comment on Neda Vishlaghi et al.: https://doi.org/10.1172/JCI201199
Address correspondence to: Benjamin Levi, Department of Surgery, University of Texas Southwestern, 6,000 Harry Hines Blvd., Dallas, Texas 75,235, USA. Phone: 214.648.9017; Email: Benjamin. [email protected].
Find articles by Vishlaghi, N. in: | Google Scholar
1Department of Surgery, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
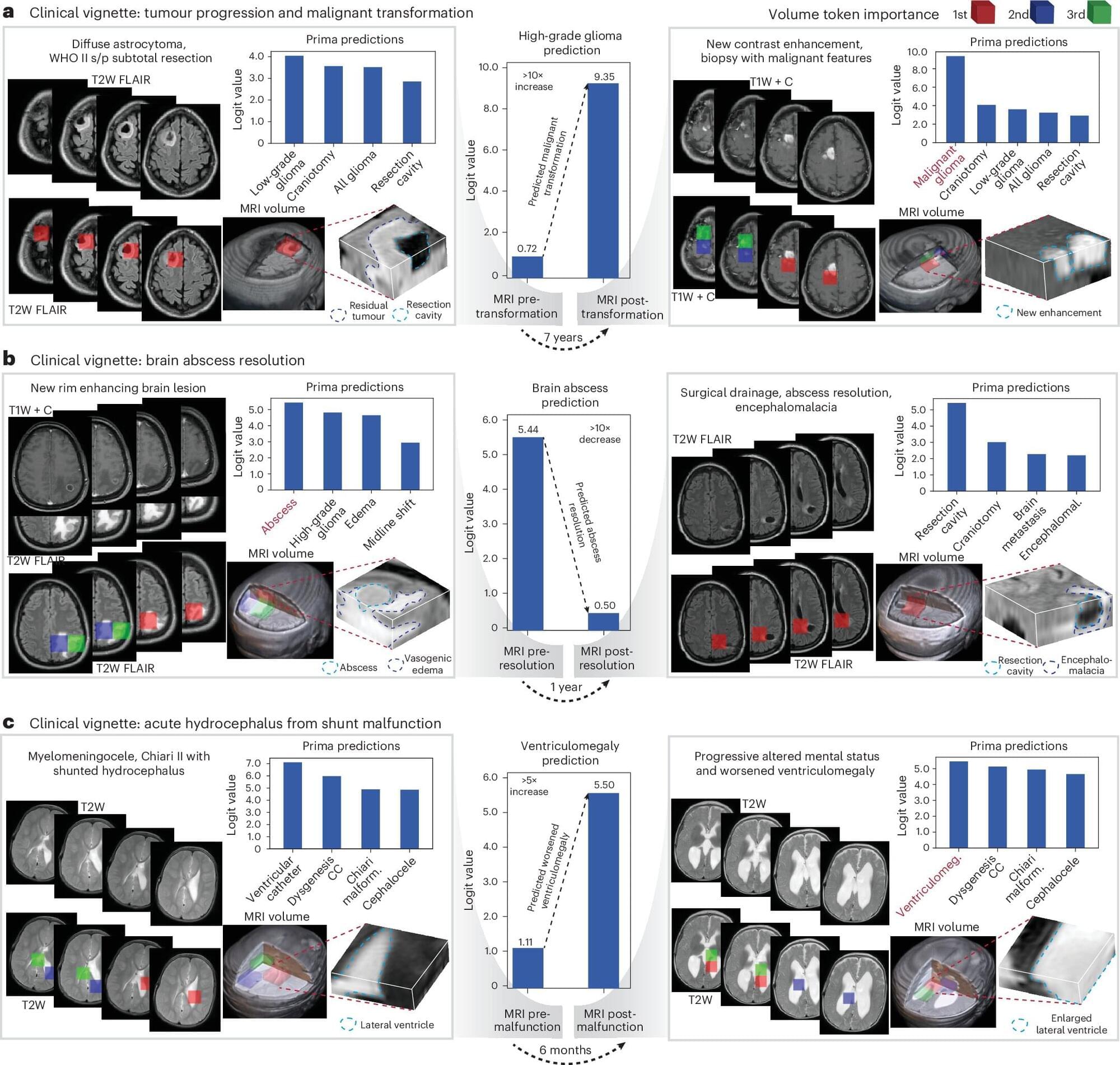
An AI-powered model developed at the University of Michigan can read a brain MRI and diagnose a person in seconds, a study suggests. The model detected neurological conditions with up to 97.5% accuracy and predicted how urgently a patient required treatment.
Researchers say the first-of-its-kind technology could transform neuroimaging at health systems across the United States. The results are published in Nature Biomedical Engineering.
“As the global demand for MRI rises and places significant strain on our physicians and health systems, our AI model has the potential to reduce burden by improving diagnosis and treatment with fast, accurate information,” said senior author Todd Hollon, M.D., a neurosurgeon at University of Michigan Health and assistant professor of neurosurgery at U-M Medical School.
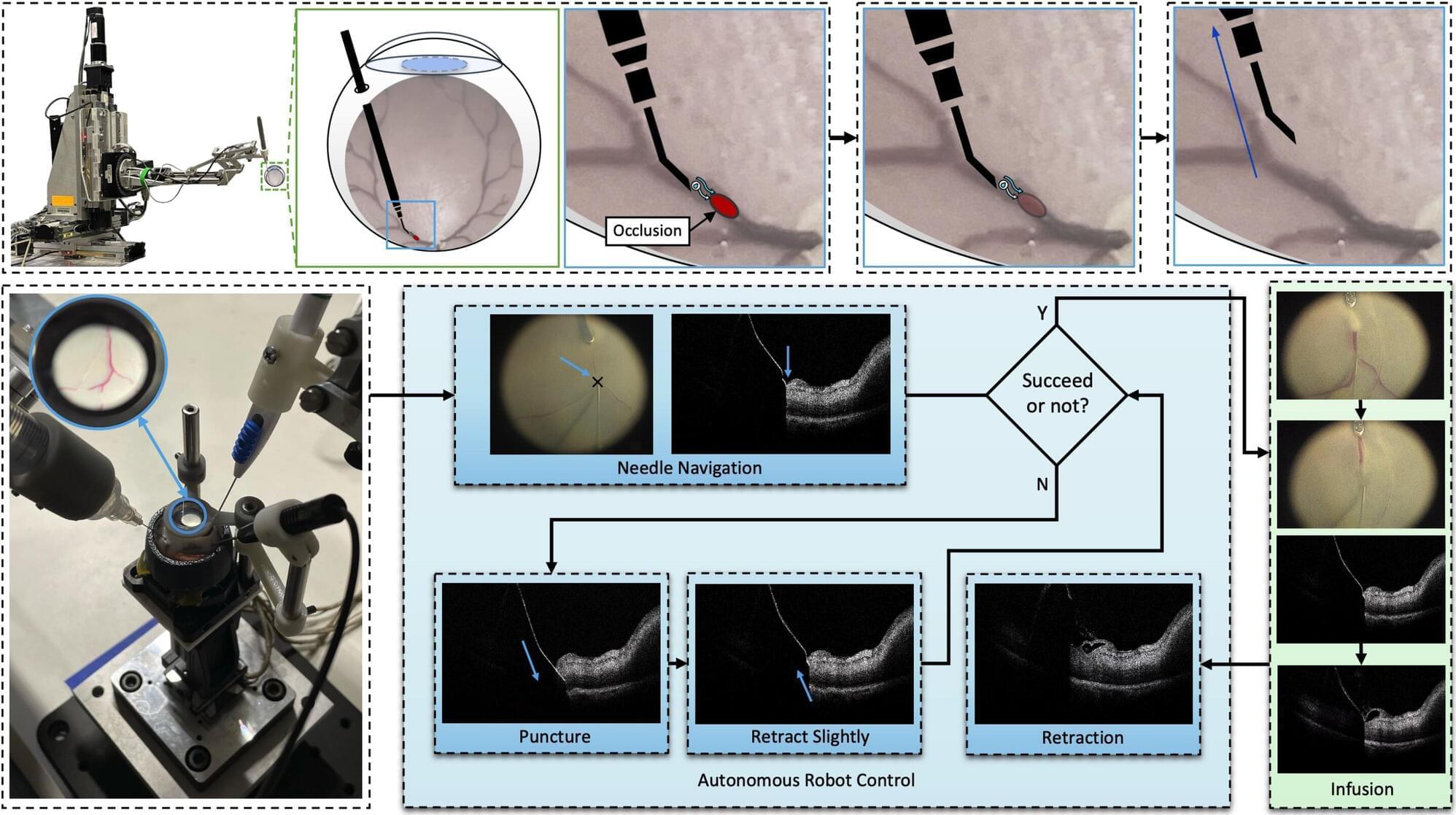
Retinal vein occlusion (RVO) is a severe disease that occurs when a vein in the light-sensitive layer at the back of the eye (i.e., the retina) becomes blocked, which results in a loss of vision. There are currently a few medical interventions that address RVO, including the periodic injection of medications that block the abnormal growth of blood vessels or of steroids, which reduce swelling and inflammation.
A promising procedure for the treatment of RVO is retinal vein cannulation (RVC). This is a very delicate surgical intervention that requires surgeons to insert a tiny needle into the blocked vein with high precision, delivering clot-dissolving drugs or medications that control the abnormal growth of blood vessels.
Given that retinal veins targeted for cannulation are similar in thickness to a human hair, manually inserting a needle inside them with high precision is very challenging. Robots could potentially assist surgeons in performing RVO procedures, ensuring that needles are inserted correctly and without damaging the patients’ retina.

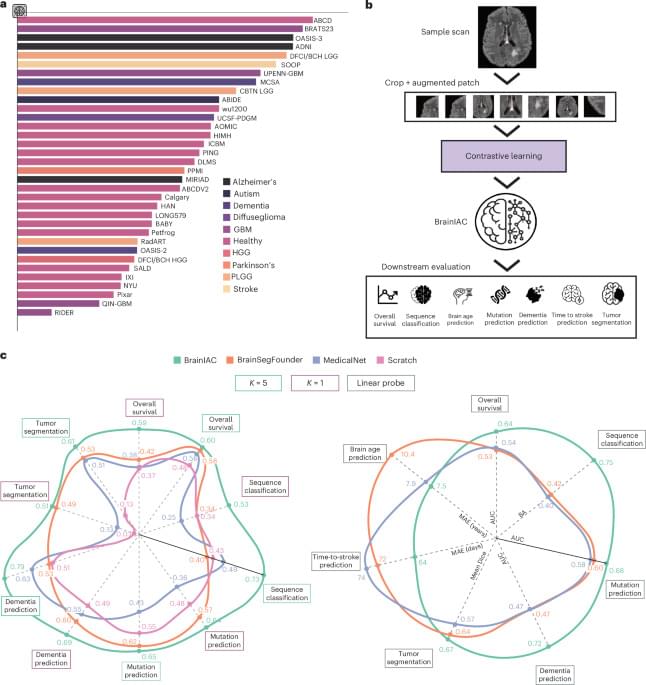
Mass General Brigham investigators have developed a robust new artificial intelligence (AI) foundation model that is capable of analyzing brain MRI datasets to perform numerous medical tasks, including identifying brain age, predicting dementia risk, detecting brain tumor mutations and predicting brain cancer survival. The tool, known as BrainIAC, outperformed other, more task-specific AI models and was especially efficient when limited training data were available.
Results are published in Nature Neuroscience.
“BrainIAC has the potential to accelerate biomarker discovery, enhance diagnostic tools and speed the adoption of AI in clinical practice,” said corresponding author Benjamin Kann, MD, of the Artificial Intelligence in Medicine (AIM) Program at Mass General Brigham. “Integrating BrainIAC into imaging protocols could help clinicians better personalize and improve patient care.”
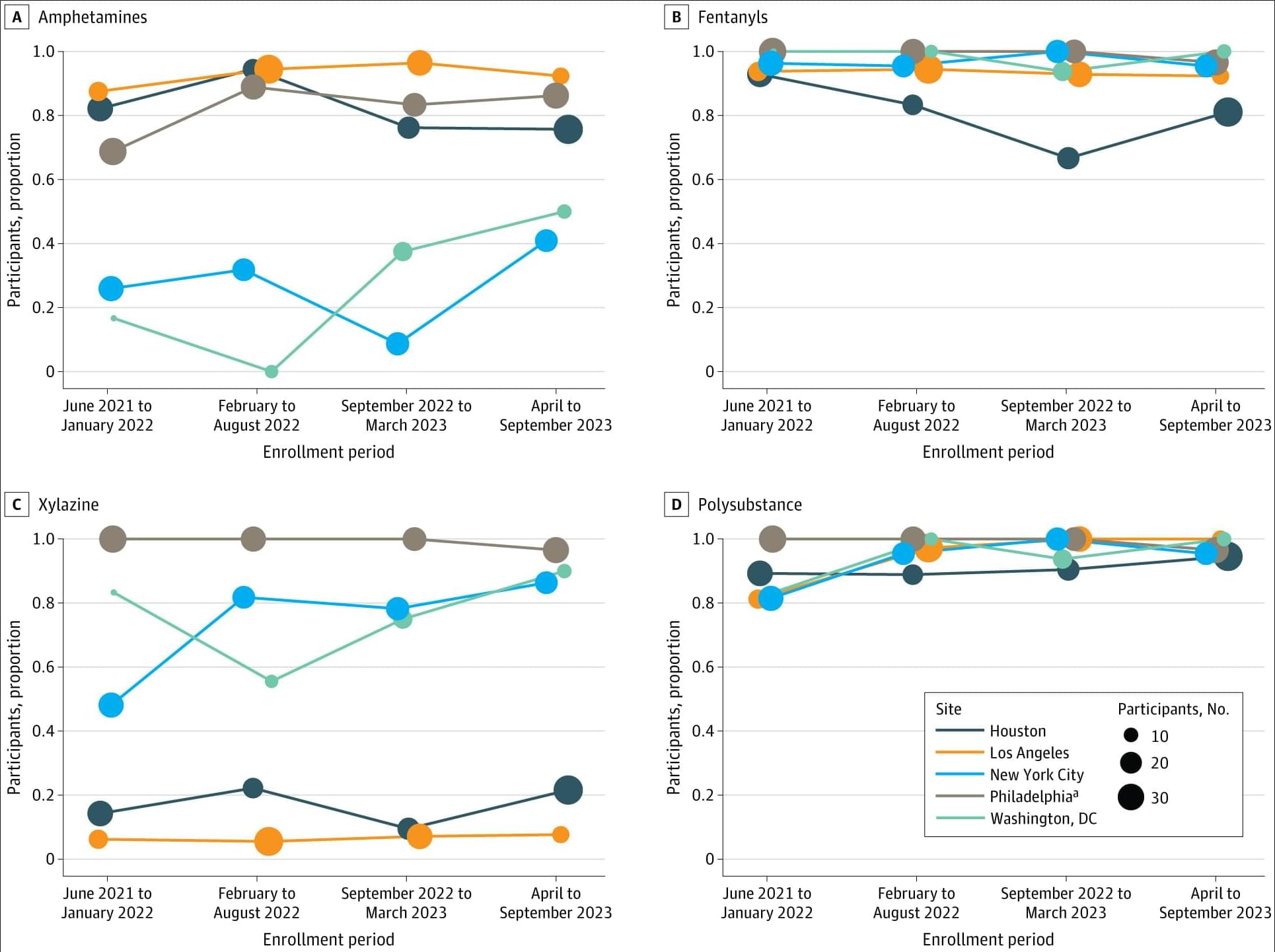
Among people injecting drugs and not engaged in medical care, nearly all tested positive for fentanyl and multiple other substances, with polysubstance and xylazine detection rates highest among unhoused and recently incarcerated participants.
This cross-sectional study used data from HPTN 094. Participants who met eligibility criteria were invited to participate in a baseline interview and were enrolled between June 2021 and September 2023. All participants completed written informed consent prior to participating in study procedures, and a single institutional review board (Advarra) provided ethical approval for HPTN 094; this cross-sectional analysis was exempt from additional IRB approval. The current study was conducted and reported in accordance with the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) reporting guideline.23
Participants were required to meet the following criteria: be at least 18 years of age, have a urine test positive for recent opioid use and evidence of recent injection drug use (visible venipuncture marks), meet diagnostic criteria for opioid use disorder, be able to give informed consent, be willing to start MOUD treatment, complete an assessment of understanding, have confirmed HIV seropositivity or self-reported sharing of injection equipment and/or condomless sex in the past 3 months with partners living with HIV or with unknown HIV status, and provide locator information. Participants were excluded if they self-reported being prescribed MOUD in the 30 days prior to screening, had a urine test positive for methadone (with the exception of verified hospitalization), or were enrolled in another study.