The genetic mutation that drives evolution is random. But here’s a list of some beneficial mutations that are known to exist in human beings.


The genetic mutation that drives evolution is random. But here’s a list of some beneficial mutations that are known to exist in human beings.

We may have parted ways with our primate cousins millions of years ago, but a new study shows just how human beings continue to evolve in ways we never imagined.
Researchers from Biomedical Sciences Research Center “Alexander Fleming” (BSRC Flemming) in Greece and Trinity College Dublin, Ireland, have identified 155 genes in our genome that emerged from small, non-coding sections of DNA. Many appear to play a critical role in our biology, revealing how completely novel genes can rapidly evolve to become essential.
New genes typically arise through well known mechanisms like duplication events, where our genetic machinery accidently produces copies of pre-existing genes that can end up suiting new functions over time.
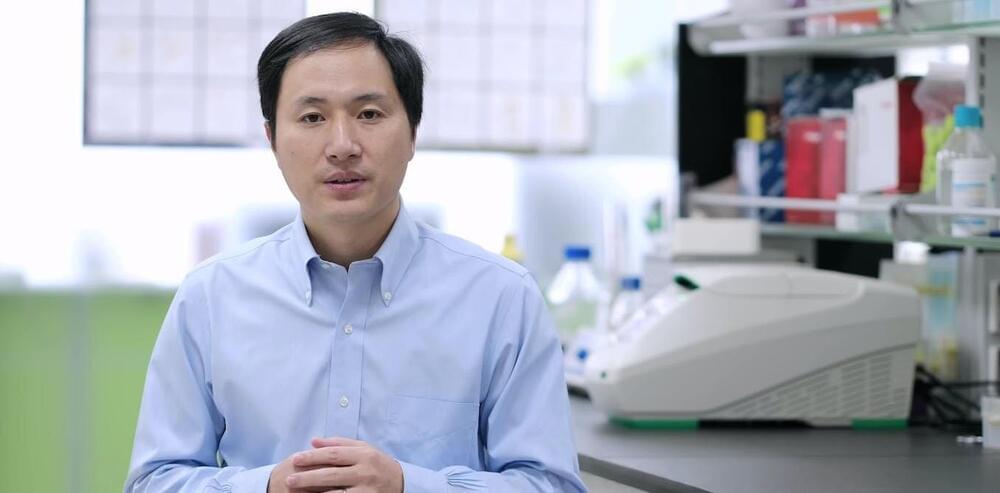

In the four years since an experiment by disgraced scientist He Jiankui resulted in the birth of the first babies with edited genes, numerous articles, books and international commissions have reflected on whether and how heritable genome editing—that is, modifying genes that will be passed on to the next generation—should proceed. They’ve reinforced an international consensus that it’s premature to proceed with heritable genome editing. Yet, concern remains that some individuals might buck that consensus and recklessly forge ahead—just as He Jiankui did.

Join us on Patreon!
https://www.patreon.com/MichaelLustgartenPhD
TruDiagnostic Discount Link (Epigenetic Testing)
CONQUERAGING!
https://bit.ly/3Rken0n.
Bristle Discount Link (Oral Microbiome Quantification):
ConquerAging15
https://www.bmq30trk.com/4FL3LK/GTSC3/
Quantify Discount Link (At-Home Blood Testing):
https://getquantify.io/mlustgarten.
Cronometer Discount Link (Daily Diet Tracking):
https://shareasale.com/r.cfm?b=1390137&u=3266601&m=61121&urllink=&afftrack=
If you’d like to support the channel, you can do that with the website.
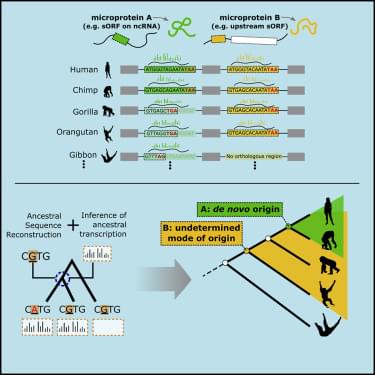
Human microproteins encoded by small ORFs have been found to be functional. By comparing the corresponding sequences across vertebrate genomes, Vakirlis et al. show that a number of these originated “from scratch” from noncoding sequences, including two very recent cases unique to humans. These cases demonstrate the rapid evolution of genetic novelty.
It will provide a better understanding of how drugs affect men and women differently.
Scientists created male and female cells with the same genetic code from the same person for the first time. This unique set of cells could provide researchers with valuable insights into how sex chromosomes affect various diseases and their role in early development.
CDC/Dr. Laine.
The sex chromosome.

This impressive achievement could potentially revolutionize how we treat cancer and immunity deficiencies.
Children born with Artemis-SCID face many challenges, from a missing repertoire of T and B cells to reduced resistance against chemotherapy used in bone marrow transplants. Additionally, malfunctioning DNA repair mechanisms increase the risk of developing graft-versus-host disease, where the donor’s immune system attacks host tissues.
That’s why researchers are trying everything to find an antidote for such a rare genetic disease and have now turned to gene therapy to treat Artemis-SCID. Gene therapy eliminates the need for donor cells.
Infant gene therapy – a breakthrough to save Artemis-SCID children
In a recent medical breakthrough, scientists have discovered how to use gene therapy to treat babies born with Severe Combined Immunodeficiency (SCID), or “bubble boy syndrome,” without needing immune-suppressing drugs.
This new innovation has proven to be potentially life-changing for infants suffering from rare diseases, giving them an exponentially improved chance of leading a relatively healthy and normal life.

The newly-created Longevity Escape Velocity Foundation (LEV) has released details of the first study in its flagship research programme: Robust Mouse Rejuvenation – Study 1.
Longevity. Technology: A highlight of Longevity Summit Dublin 2022 was Dr Aubrey de Grey’s announcement of his new foundation; LEV Foundation exists to proactively identify and address the most challenging obstacles on the path to the widespread availability of genuinely effective treatments to prevent and reverse human age-related disease, and to that end, its flagship research programme is a sequence of large mouse lifespan studies.
Mouse models are significant in aging research for several reasons. Mice and humans share many genetic and physiological similarities, including similar aging-related pathways, and this makes mice a useful model for studying the molecular and cellular processes underlying aging in humans.