Among survivors of non-small cell lung cancer, nearly 8% developed non-lung secondary cancers over about 6 years, and this risk may be driven by genetic factors.


Researchers say they have created the world’s first scalable atomic quantum processor that achieves record-breaking 99.99% fidelity.
Elon Musk is considering Tesla taking a pre-IPO stake in SpaceX to integrate their businesses, accelerate ambitious projects, and increase the value of both companies ## ## Questions to inspire discussion.
Strategic Governance Alignment.
🔄 Q: Why should Tesla acquire a pre-IPO stake in SpaceX rather than waiting until after the IPO? A: A pre-IPO stake resolves governance and conflict risks before SpaceX’s planned $30B IPO in mid-2026, ensuring all transactions are recorded as part of the IPO and avoiding complications that could impact IPO pricing or create persistent post-IPO conflicts between the two companies.
🎯 Q: What is the core governance problem Tesla shareholders currently face with SpaceX? A: Tesla shareholders are exposed to SpaceX outcomes through dependencies on Starlink connectivity, orbital compute, and launch cadence without any ownership rights, governance rights, or downside protection as the companies converge operationally but not financially.
⚖️ Q: How would a pre-IPO stake transaction affect Tesla’s ownership structure and Musk’s control? A: The transaction would dilute Tesla by 20% but could raise market cap to $1.62-2T, increasing Musk’s stake to 22.1–24% and his net worth approaching $1T, enabling him to achieve 25% control significantly earlier than under the compensation plan.
Capital Requirements and Infrastructure.
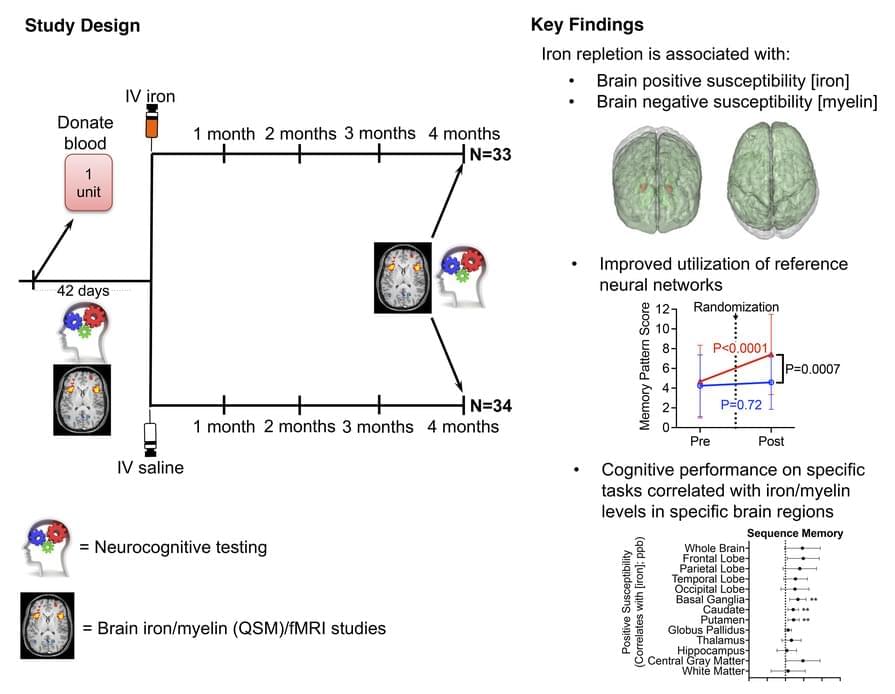
Steven L. Spitalnik & team report on a double-blind randomized trial for iron-deficient blood donors, finding treatment appears to affect brain function, brain iron, and myelin levels:
The heatmap images highlight the trend for increased iron in most brain regions.
1Department of Pathology and Cell Biology, and.
2Cognitive Neuroscience Division in Neurology, Columbia University College of Physicians and Surgeons, New York Presbyterian Hospital, New York, New York, USA.
3Department of Radiology, Weill Cornell Medical College, New York, New York, USA.
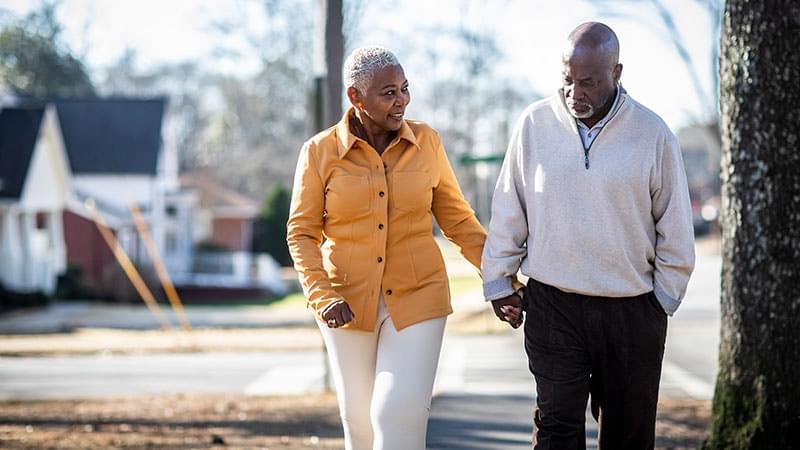

Adults with chronic heart failure with reduced ejection fraction (HFrEF) who wore a lightweight exosuit during exercise showed significant improvements in the 6-minute walk distance and daily step counts, which were not statistically significant in those who received nonassisted conventional exercise training.
Robot-assisted training with a lightweight exosuit may help patients with advanced heart failure walk more and help them stay engaged in rehabilitation, a study finds.
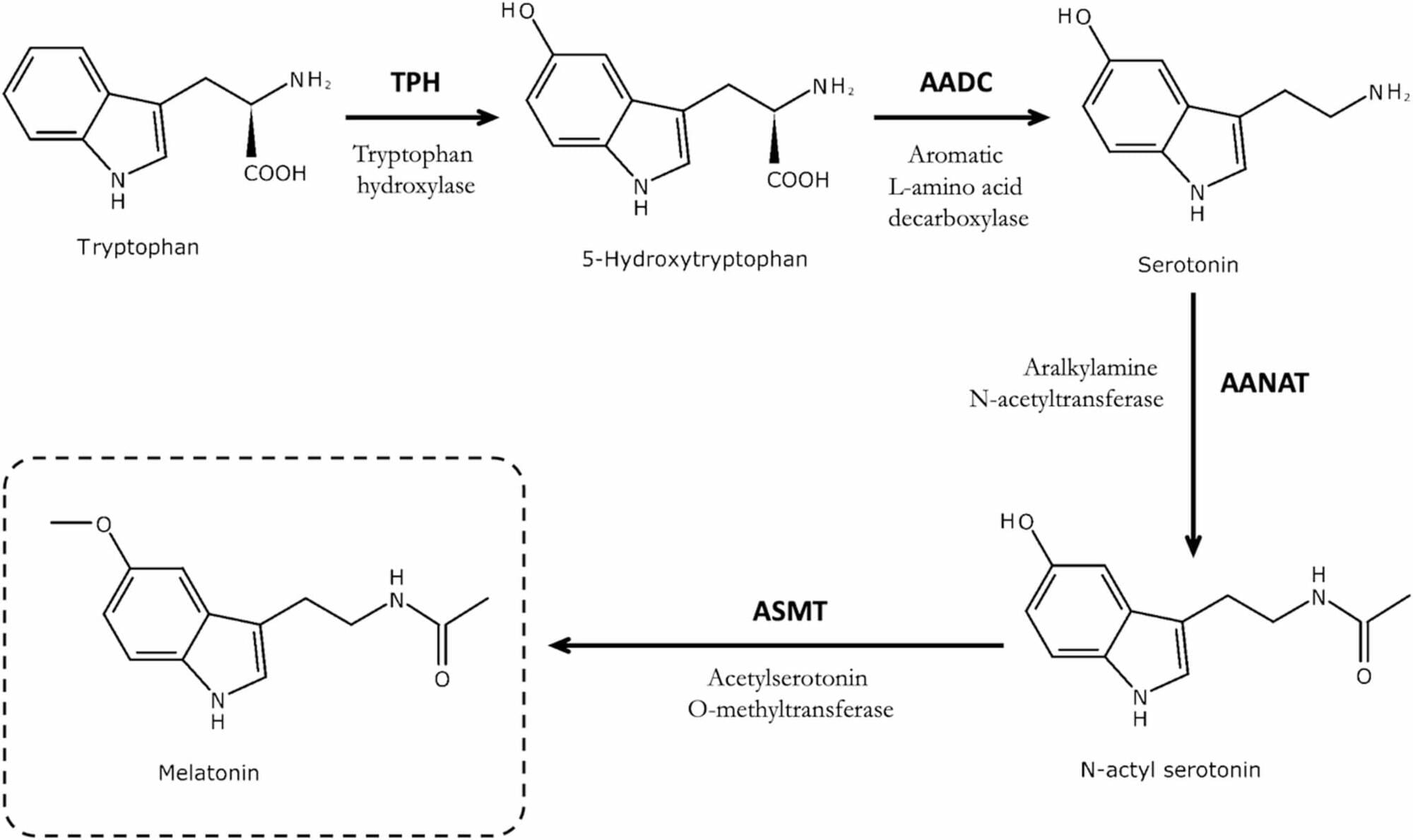
An ageing-related pathology has recently been described as one that develops and/or progresses with increasing chronological age, that is associated with, or contributes to, functional decline and that is evidenced by studies in humans. The pineal gland is a photo-neuroendocrine organ whose primary function is to produce and secrete melatonin in response to light-dark cycle environmental cues. The gland may undergo ageing-related structural and morphological changes, including calcification, gliosis, cyst formation, and reduced density of β-adrenergic receptors, which are hypothesised to reduce melatonin secretion.
The breakthrough wasn’t speed or scale, but a neural architecture that finally respected protein geometry.
AlphaFold didn’t accelerate biology by running faster experiments. It changed the engineering assumptions behind protein structure prediction.