The U.S. Food and Drug Administration (FDA) has granted orphan drug designation to a new gene therapy for Amyotrophic Lateral Sclerosis (ALS) developed at the Universitat Autònoma de Barcelona and licensed to the U.S. company Klotho Neurosciences, Inc.
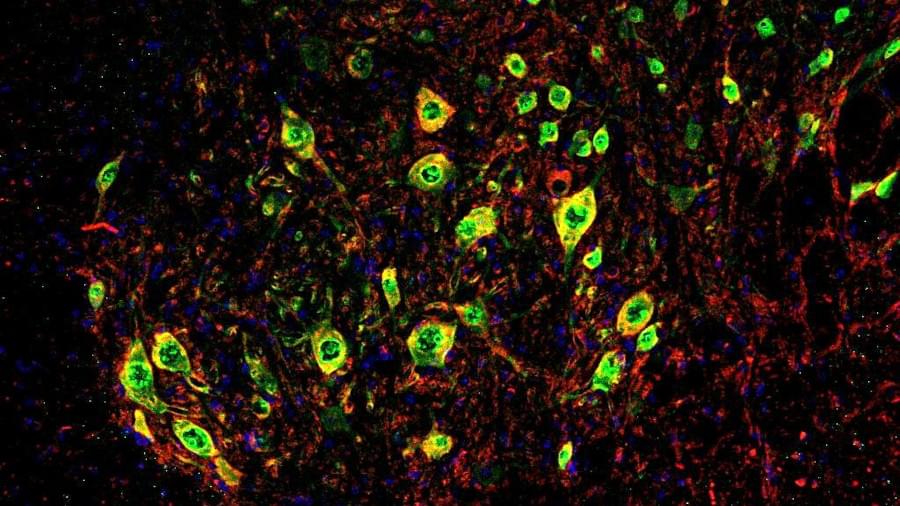
The drug uses a viral vector of the AAV (adeno-associated virus) type that expresses the secreted isoform of Klotho (s-KL) protein, with neuroregenerating, antioxidant and anti-inflammatory properties. In order to reach the neuromuscular junctions affected by the ALS disease, the vector acts under the control of a DNA sequence that regulates the expression of the protein specifically in the muscle (a muscle-specific promoter), so that therapeutic activity is directed towards the neuromuscular junctions. This innovative approach has shown very promising results in the most widely used mouse model for the preclinical study of ALS, delaying the onset of the disease, preserving neuromuscular function and extending survival.
The technological development was led by UAB researchers, with the involvement of the CIBER, ICREA and Vall d’Hebron Research Institute, co-owners of the intellectual property relating to the use of the Klotho protein and licensed to Klotho Neurosciences –a start-up company based on knowledge generated at UAB and listed on Nasdaq in 2023 (NASDAQ: KLTO)-. The technology was developed by the research groups of Assumpció Bosch and Miquel Chillón, both from the UAB Department of Biochemistry and Molecular Biology and the UAB Institut de Neurociències (INc-UAB). The research project also included the collaboration of the group led by Professor Xavier Navarro, researcher at the Institut de Neurociències and the UAB Department of Cellular Biology, Physiology and Immunology, and expert in neuroregeneration and motor neuron diseases.
“The orphan drug designation for the therapy we have developed acknowledges the relevance of treatments targeting muscle and neuromuscular junction as a strategy for ALS”, says Assumpció Bosch, principal investigator of the study. “To date, we have been able to demonstrate efficacy in a leading animal model for this pathology. We are now testing it in other ALS models to confirm that this therapeutic solution can be applied to the widest possible number of patients”, adds Sergi Verdés, postdoctoral researcher on the research team.
Receiving the orphan drug designation by the FDA underscores the potential of the treatment for the rare and severely disabling disease ALS, which affects around 65,000 people in Europe and for which there is no effective treatment. This recognition offers advantages such as seven years of exclusivity for the drug in the U.S. market, fee waivers and tax incentives for clinical trials.