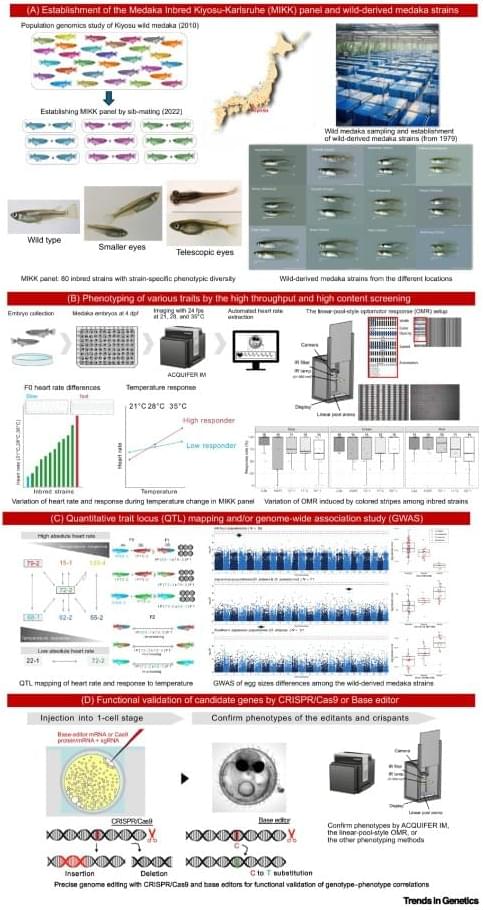
Analyzing genome– environment interactions using medaka model.
As a temperate fish species, medaka tolerates fluctuating environments (e.g., seasonal changes and different water salinity levels).
Medaka are highly tolerant to inbreeding; a near-isogenic inbred panel named Medaka Inbred Kiyosu-Karlsruhe panel and several wild-derived inbred strains of medaka have been established.
These panels and strains allow the analysis of phenotype–genotype interactions under different environmental conditions and the modeling of human populations.
Advanced epigenomic approaches, including assay for transposase-accessible chromatin using sequencing, highthroughput chromosome conformation capture (Hi-C), and analysis of histone modifications and DNA methylation, extend genomic approaches in our understanding of gene–environment interactions. sciencenewshighlights ScienceMission https://sciencemission.com/Medaka
Medaka is an established vertebrate model system for biological and biomedical research. It possesses unique features that make it particularly suitable for studying genome–environment interactions. Endemic to habitats spanning from 4 to 40°C and varying salinities, it combines broad ecological adaptability with experimental tractability. Its exceptional tolerance to inbreeding enabled the creation of the Medaka Inbred Kiyosu-Karlsruhe panel—80 near-isogenic, fully sequenced lines derived from a single wild population. More than 100 wild-derived, fully sequenced strains, collected throughout East Asia for more than 40 years, show relatively low intra-strain variation (inbreeding coefficient of 0.75) but high inter-strain variability (SNP rates 4%). Advanced quantification methods facilitate genome-wide association studies and quantitative trait locus mapping.