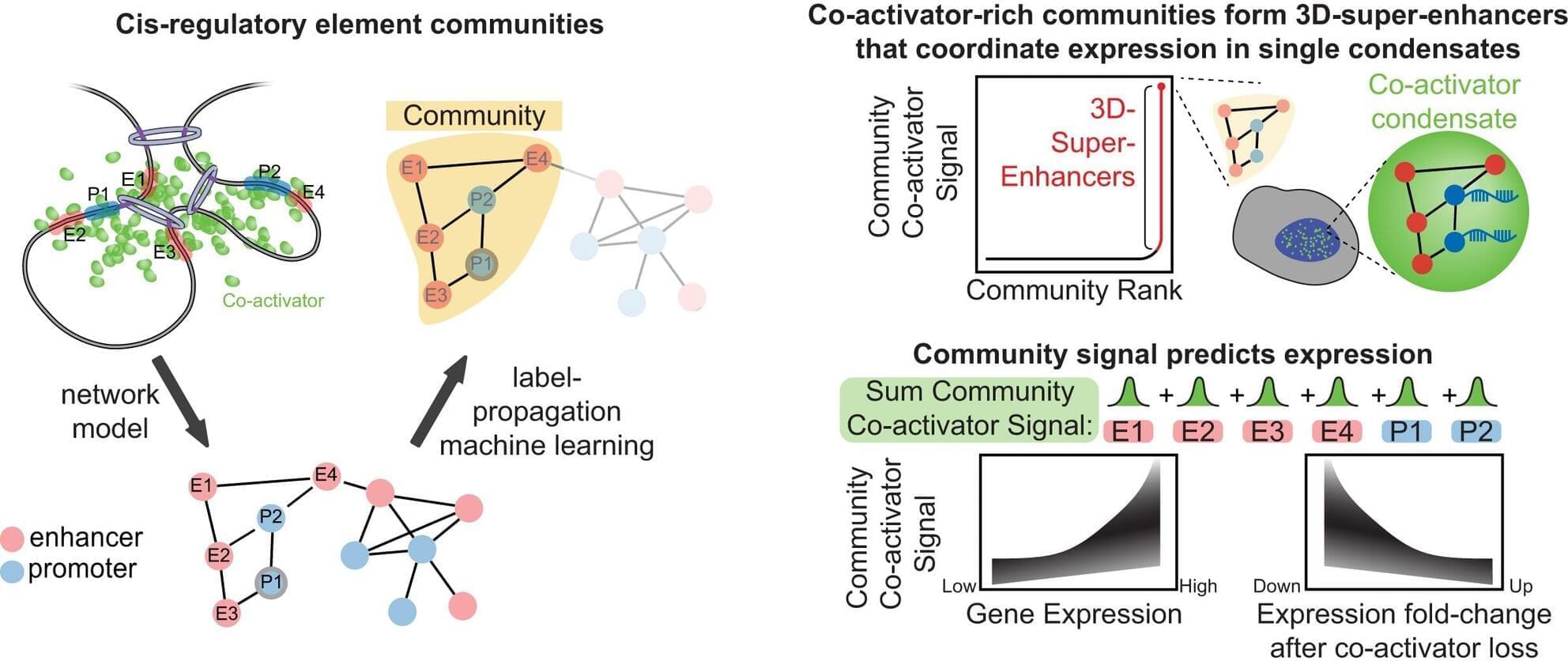
Scientists usually study the molecular machinery that controls gene expression from the perspective of a linear, two-dimensional genome—even though DNA and its bound proteins function in three dimensions (3D). To better understand how key components of this machinery, such as super-enhancers, regulate genes in this 3D reality, scientists at St. Jude Children’s Research Hospital have developed a new algorithm called BOUQUET.
Using machine learning, BOUQUET reveals that sets of genes and their regulatory elements can interact within protein condensates, high-density membraneless droplets, in cells’ nuclei. The findings, which provide new insight into how cells regulate the genes that control their specialized identities, were published today in Nucleic Acids Research.
Cells express certain sets of genes to carry out specific functions; for example, a blood cell and a brain cell express different context-specific genes. There are 3 billion base pairs of human DNA, and the genes involved in cell identity are scattered throughout. Even more challenging, enhancers, DNA elements that activate gene expression, can be thousands of DNA bases away from their target genes.