The researchers first gave a bout of psoriasis to mice when they were young. They discovered that about 10–15% of the memories that persisted a month later stuck around even to the end of the mouse’s life (~2 years). To see why these long-term memories lingered while their short-term counterparts faded within six months, they analyzed the DNA sequence characteristics within each of the memories by using a deep learning model customized by the third co-first author.
“When we compared the DNA sequences of short and long-term memory domains, they looked very similar in terms of the numbers and kinds of transcription factor binding sites,” says the author. “We realized we needed to develop a new metric that specifically captures memory persistence across time, not just total accessibility at any one point.”
Soto-Ugaldi’s adaptation, called PersistNet, quickly identified a telling trait: The longest lasting memory domains had an unusually high frequency of CpG dinucleotides—short DNA sequences of cytosine followed by guanine, which are known to play a key role in gene regulation. In fact, the model predicted that CpG density hardwires a timer into every memory domain: The more CpG’s, the longer the memory.
When they tested the prediction, that’s exactly what they found. “Looking across all 1,000 memory domains, we discovered that these nucleotide densities alone, and no other DNA sequence pattern, could distinguish how long each memory would linger,” says the author.
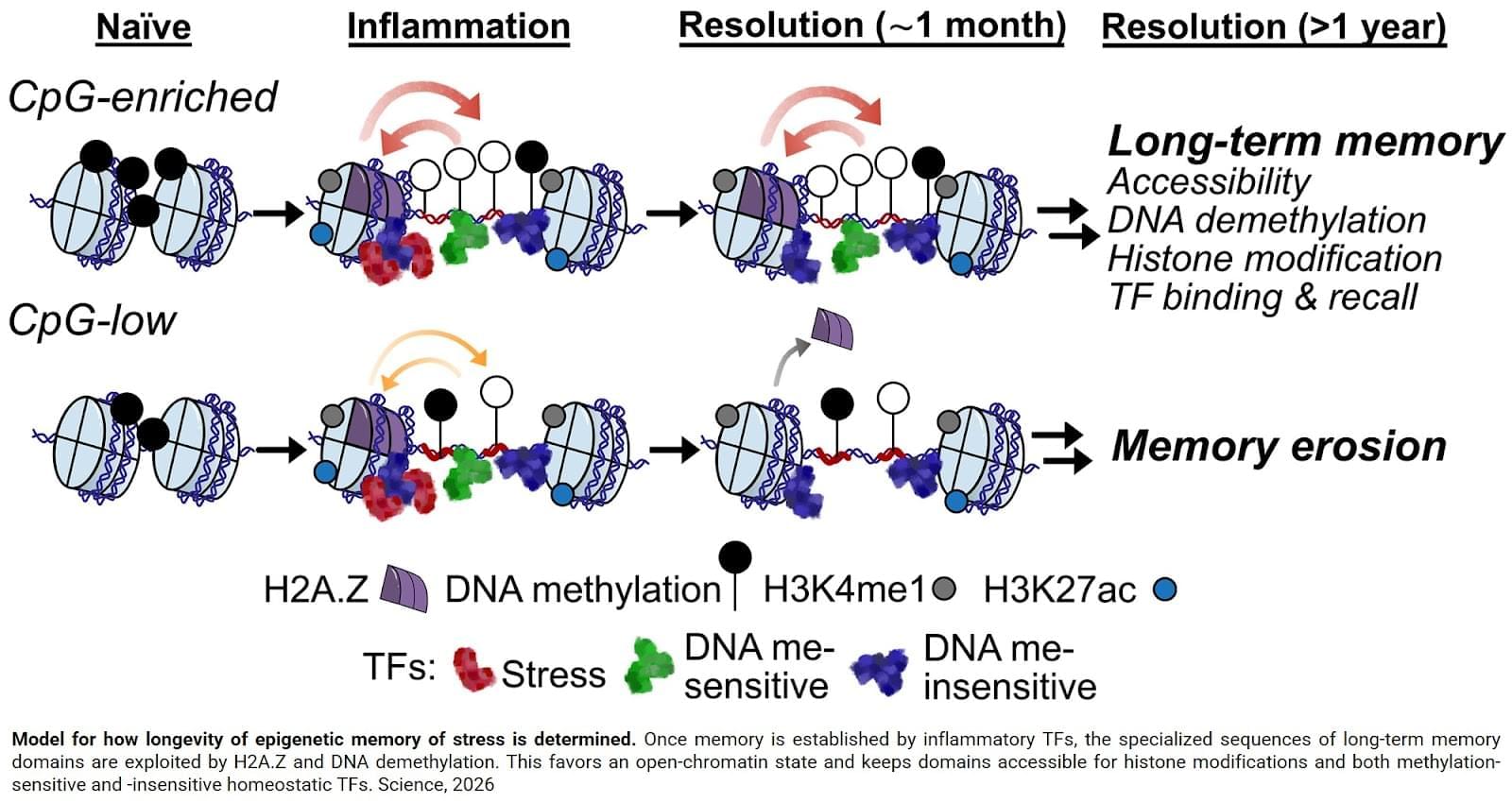
Back in the lab, the team discovered that these genetically wired densities enabled a host of epigenetic changes in memory domains, including DNA demethylation (the removal of a methyl group specifically found on CpG dinucleotides); the binding of transcription factors that prefer demethylated states; and the recruitment of a histone variant called H2A.Z, which preferentially seeks out demethylated sites and boosts chromatin accessibility while staving off future re-methylation. Together, these changes stabilized the open chromatin formation and its gene-priming activity. As the authors discovered, this structure could crucially be passed down across cellular generations, essentially keeping the doors open for life. Science Mission sciencenewshighlights.
One of the most puzzling aspects of common chronic inflammatory skin diseases such as psoriasis is how they become chronic. What allows an ongoing condition to stay dormant for months or even years, then seemingly spring back out of nowhere?
In 2017, a research team tackled one piece of that puzzle by investigating a curious phenomenon: Flareups often recur in the same locations. In the process, they discovered that our skin’s stem cells can retain “memories” of past experiences in a way that (in healthy responses) prepares them to heal injuries faster in the future. If these responses become dysfunctional, however, “bad” memories can make the tissue hypersensitive to environmental triggers and lead to chronic inflammation, as in psoriasis.
Whether good or bad, it remained a mystery as to how memories of past inflammation can persist for years. Now the team has identified tantalizing clues. In a new paper in Science, they identify key new epigenetic drivers of memory persistence. Using a combination of wet lab research and deep machine learning, they pinpoint distinct genetic sequences that propel a handful of critical memories into the years-long timeframes that underpin chronic disease.