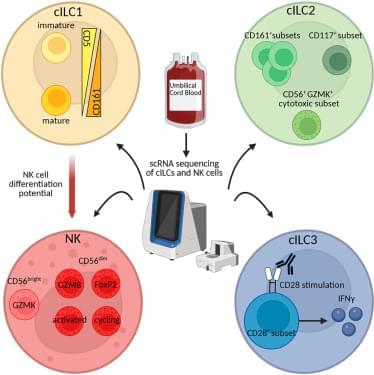
Mononuclear cells (MNCs) were isolated from fresh umbilical cord blood (CB) via a Ficoll gradient (n = 3). Unwanted cells were depleted using biotinylated antibodies (anti-CD3, anti-CD14, anti-CD19, and CD66b) and magnetic beads. The cells were stained to faithfully sort cILC1s (Lin−CD94−CD127+CD117−CRTH2−), cILC2s (Lin−CD94−CD127+CD117−/+ CRTH2+), cILC3s (Lin−CD94−CD127+CD117+CRTH2−), and NK cells (Lin−CD94+), as previously described.5,7,33 The four individual populations were individually multiplexed, pooled, and stained with Ab-Seq antibodies.
(A) scRNA-seq was performed via the BD Rhapsody protocol.
(B and C) UMAP visualization of sorted populations (B) and individual clusters of CB cILCs and NK cells with color coding of the individual cluster 0–10. The following clusters were identified: 0, CD56dim NK cells (GZMBhigh); 1, CD56dim NK cells (GZMBlow); 2, cILC2s (GATA3); 3, cILC progenitor (KIT); 4, CD56bright NK cells (GZMK); 5, intermediate zone (NK/cILCs); 6, cILC1s (CD5); 7, activated CD56dim NK cells (PCNA); 8, cycling CD56dim NK cells (MKI67); 9, cILC3s (RORC); and 10, CD56dim NK cells (FOXP2) ©.