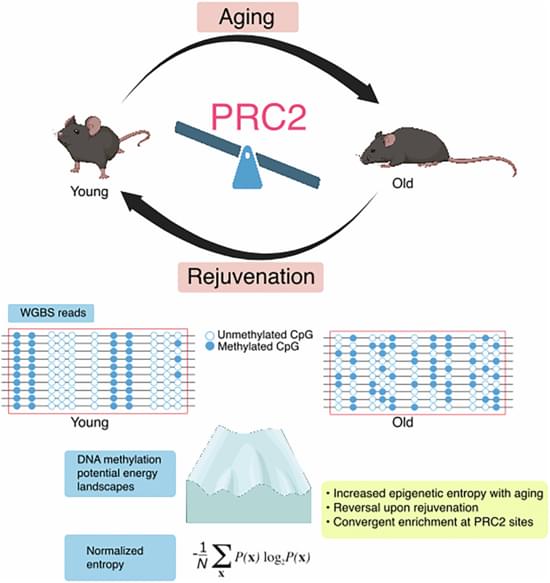
Rejuvenation of tissues in physiologically aging mice can be accomplished by long-term partial reprogramming via expression of reprogramming factors (Oct4, Sox2, Klf4, and c-Myc). To investigate the epigenetic determinants of partial reprogramming-mediated rejuvenation, we used whole-genome bisulfite sequencing to carry out unbiased comprehensive profiling of DNA methylation changes in skin from mice subjected to partial reprogramming, as well as young and untreated old controls. We found a striking convergence of age- and rejuvenation-related epigenetic alterations on targets of the Polycomb repressive complex 2 (PRC2), with increased DNA methylation level and entropy over these regions. Native ChIP demonstrated extensive loss of H3K27me3 in aged epidermis compared to young, partially overlapping regions with age- and rejuvenation-related DNA methylation changes. In addition, large H3K9me2-marked “LOCK” heterochromatin domains defined the boundaries for hypomethylated highly entropic regions during aging. These results are also supported by a likewise prominent enrichment of PRC2 targets in gene expression data, suggesting that PRC2 activity can modulate aging and mediate tissue rejuvenation.