DNA methylation has shown great potential in Alzheimer’s disease (AD) blood diagnosis. However, the ability of long non-coding RNAs (lncRNAs), which can be modified by DNA methylation, to serve as noninvasive biomarkers for AD diagnosis remains unclear.
We performed logistic regression analysis of DNA methylation data from the blood of patients with AD compared and normal controls to identify epigenetically regulated (ER) lncRNAs. Through five machine learning algorithms, we prioritized ER lncRNAs associated with AD diagnosis. An AD blood diagnosis model was constructed based on lncRNA methylation in Australian Imaging, Biomarkers, and Lifestyle (AIBL) subject and verified in two large blood-based studies, the European collaboration for the discovery of novel biomarkers for Alzheimer’s disease (AddNeuroMed) and the Alzheimer’s Disease Neuroimaging Initiative (ADNI). In addition, the potential biological functions and clinical associations of lncRNAs were explored, and their neuropathological roles in AD brain tissue were estimated via cross-tissue analysis.
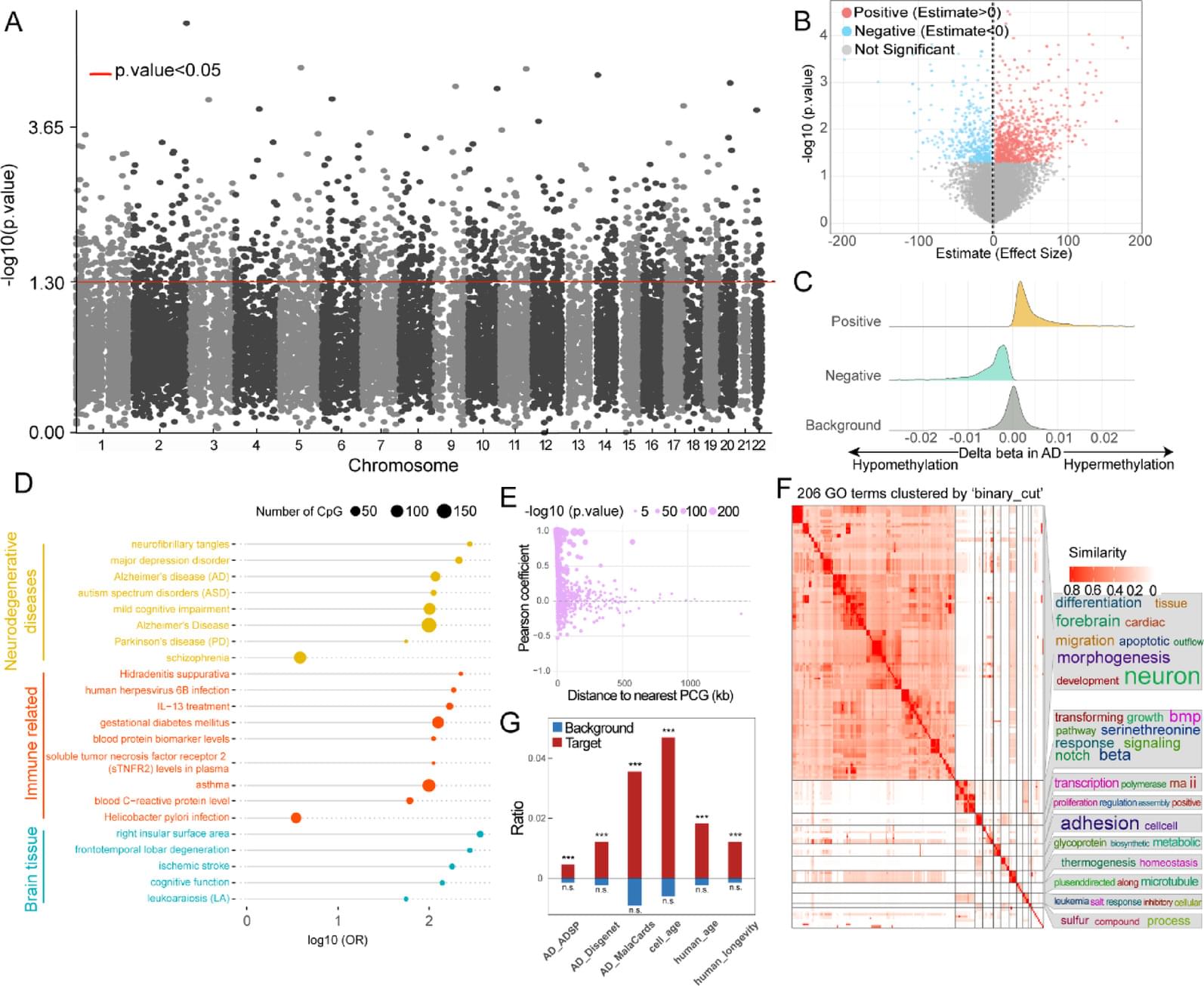
We characterized the ER lncRNA landscape in AD blood, which is strongly related to AD occurrence and process. Fifteen ER lncRNAs were prioritized to construct an AD blood diagnostic and nomogram model. The receiver operating characteristic (ROC) curve and the decision and calibration curves show that the model has good prediction performance. We found that the targets and lncRNAs were correlated with AD clinical features. Moreover, cross-tissue analysis revealed that the lncRNA ENSG0000029584 plays both diagnostic and neuropathological roles in AD.