Numerous efforts have been made to improve the temporal resolution of CRISPR-Cas9–mediated DNA cleavage to the hour time scale. Liu et al. developed a Cas9 system that achieved genome-editing manipulation at the second time scale (see the Perspective by Medhi and Jasin). Part of the guide RNA is chemically caged, allowing the Cas9-guide RNA complex to bind at a specific genomic locus without cleavage until activation by light. This fast CRISPR system achieves genome editing at high temporal resolution, enabling the study of early molecular events of DNA repair processes. This system also has high spatial resolution at short time scales, allowing editing of one genomic allele while leaving the other unperturbed.
Science, this issue p. 1265; see also p. 1180
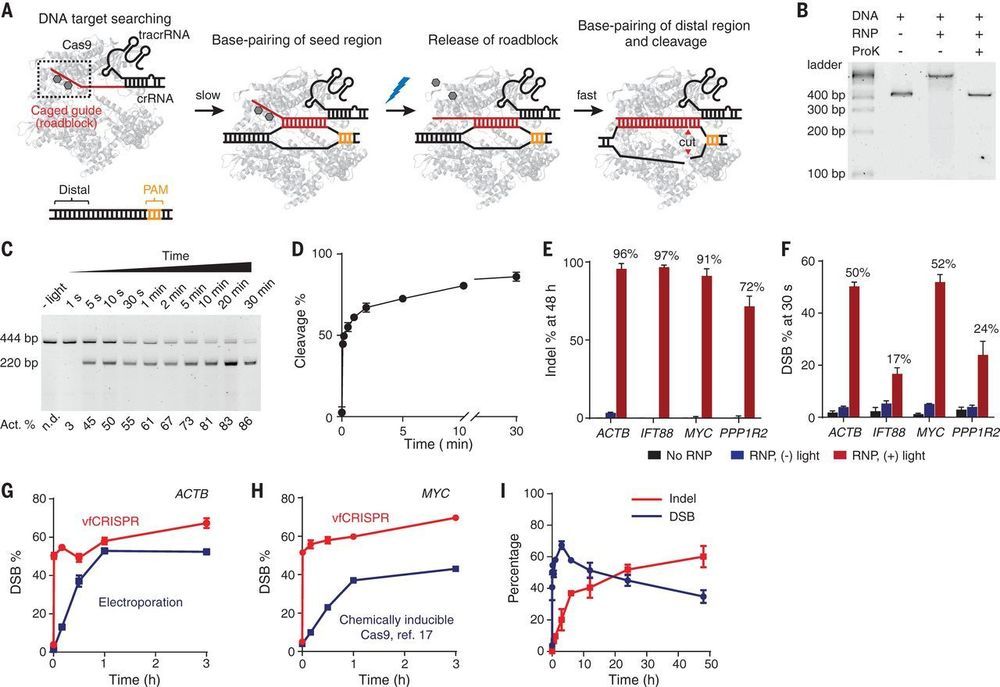
CRISPR-Cas systems provide versatile tools for programmable genome editing. Here, we developed a caged RNA strategy that allows Cas9 to bind DNA but not cleave until light-induced activation. This approach, referred to as very fast CRISPR (vfCRISPR), creates double-strand breaks (DSBs) at the submicrometer and second scales. Synchronized cleavage improved kinetic analysis of DNA repair, revealing that cells respond to Cas9-induced DSBs within minutes and can retain MRE11 after DNA ligation. Phosphorylation of H2AX after DNA damage propagated more than 100 kilobases per minute, reaching up to 30 megabases. Using single-cell fluorescence imaging, we characterized multiple cycles of 53BP1 repair foci formation and dissolution, with the first cycle taking longer than subsequent cycles and its duration modulated by inhibition of repair. Imaging-guided subcellular Cas9 activation further facilitated genomic manipulation with single-allele resolution.