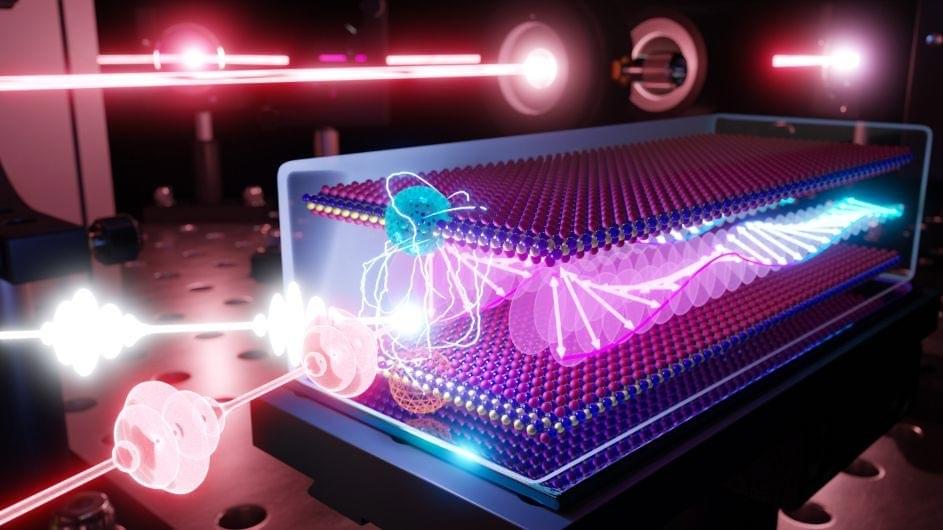
New research reveals that spinning quasiparticles, or magnons, light up when paired with a light-emitting quasiparticle, or exciton, with potential quantum information applications.
All magnets contain spinning quasiparticles called magnons. This is true of all magnets from the simple souvenirs hanging on your refrigerator to the discs that give your computer memory storage to the powerful versions used in research labs. The direction one magnon spins can influence that of its neighbor, which in turn affects the spin of its neighbor, and so on, yielding what are known as spin waves. Spin waves can potentially transmit information more efficiently than electricity, and magnons can serve as “quantum interconnects” that “glue” quantum bits together into powerful computers.
Although magnons have enormous potential, they are often difficult to detect without bulky pieces of lab equipment. According to Columbia researcher Xiaoyang Zhu, such setups are fine for conducting experiments, but not for developing devices, such as magnonic devices and so-called spintronics. However, seeing magnons can be made much simpler with the right material: a magnetic semiconductor called chromium sulfide bromide (CrSBr) that can be peeled into atom.