A new computational method could dramatically accelerate efforts to map the body’s cells in space, according to a study published in Nature Genetics. Spatial multi-omics technologies—often described as ultra-high-resolution maps of tissues—allow scientists to see not only which genes or proteins are active in a cell, but exactly where that activity occurs. That spatial context is critical for understanding complex organs such as the brain, immune tissues and developing embryos.
Unfortunately, capturing multiple molecular layers at once remains expensive and technically challenging, said David Gate, Ph.D., assistant professor in the Ken and Ruth Davee Department of Neurology’s Division of Behavioral Neurology, who was a co-author of the study.
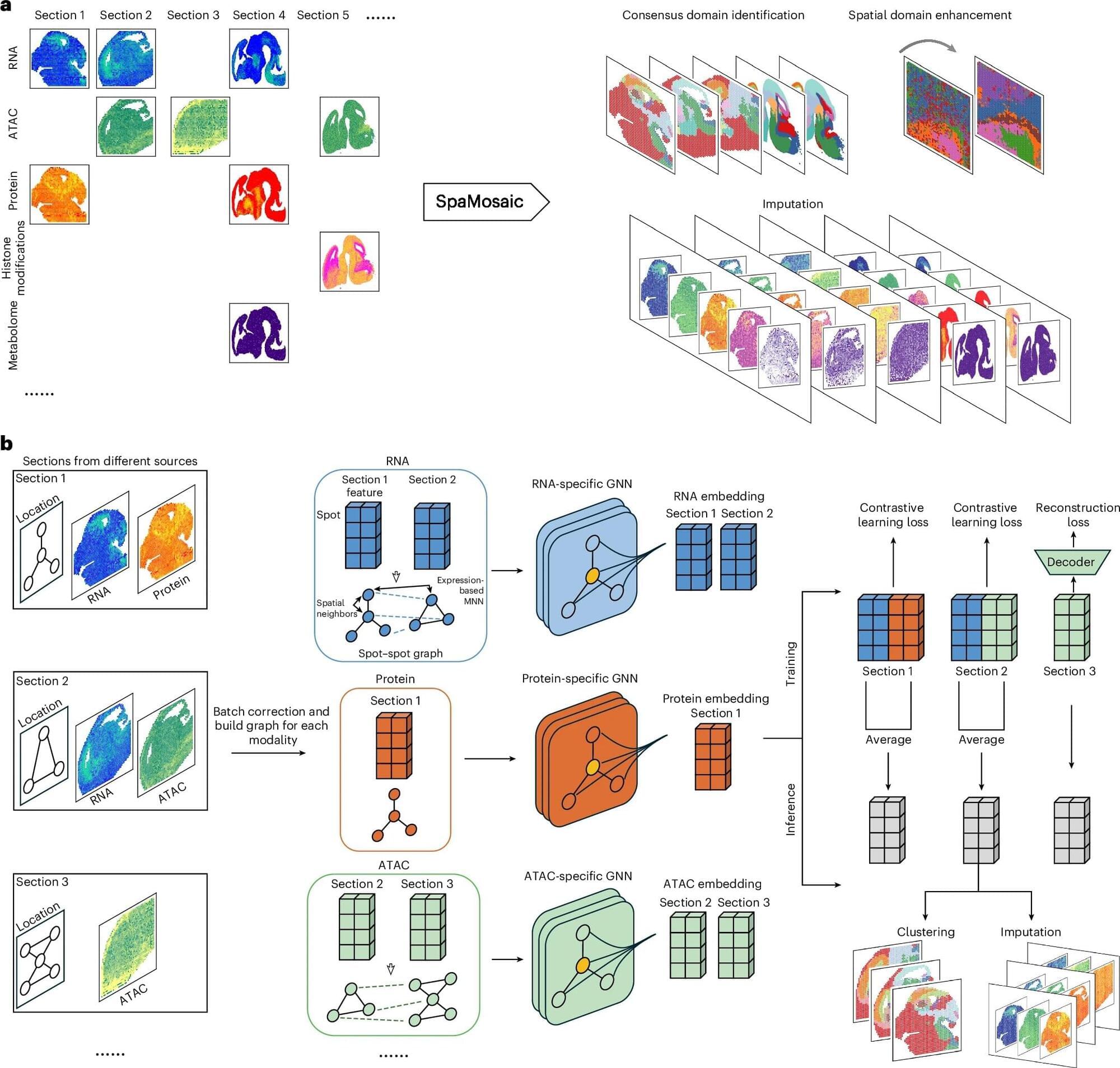
“In practice, investigators end up with ‘mosaic’ datasets: different slices or batches that each capture only some of the layers, often from different technologies or labs, with batch effects and missing pieces,” said Gate, who also leads the Abrams Research Center on Neurogenomics.