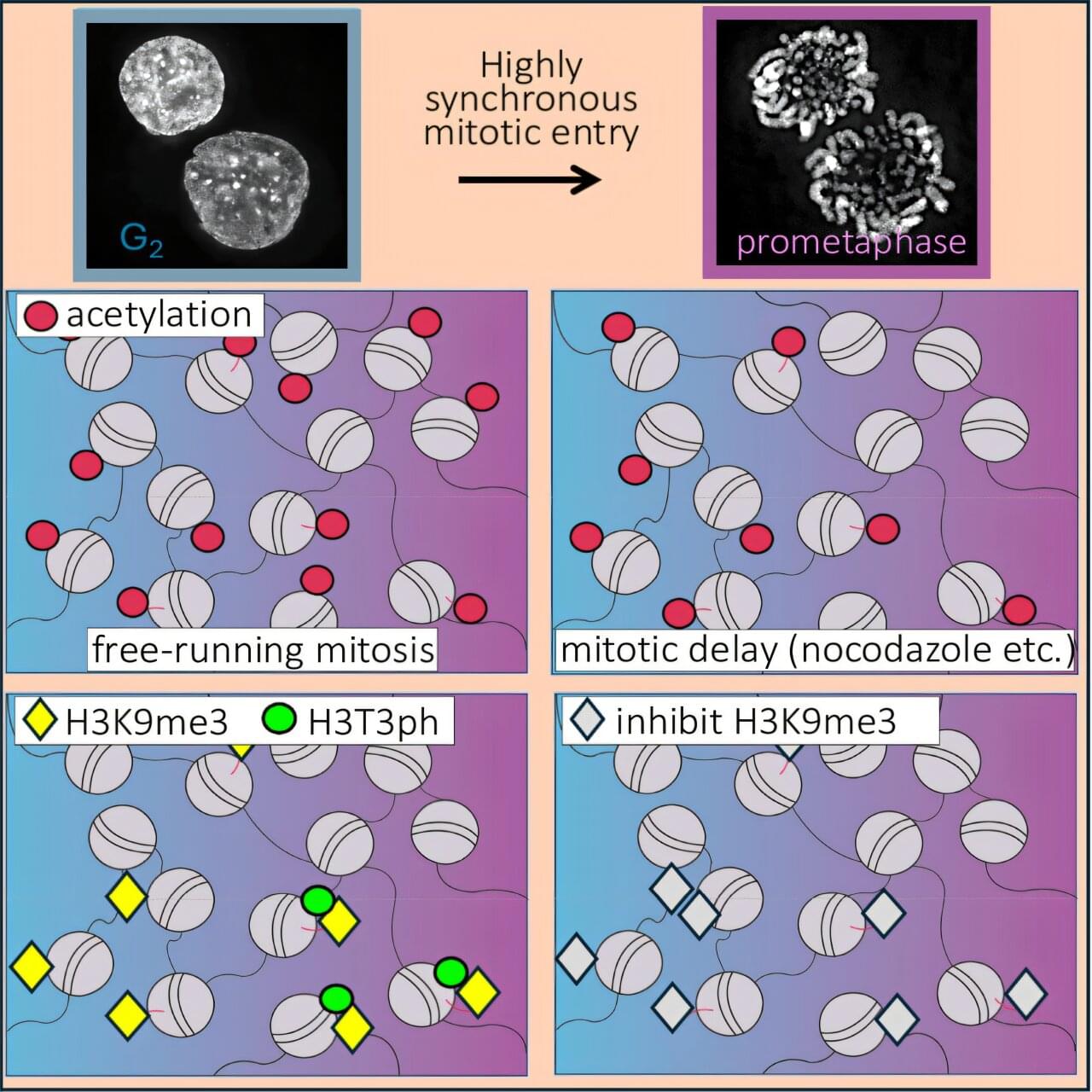
DNA does not float freely in the cell. Instead, it is wrapped around histone proteins to form structures called nucleosomes. These histones carry numerous chemical modifications that act as molecular signals, controlling how tightly the DNA is packaged and which genes are active. During cell division, this DNA-histone complex—known as chromatin—must be further condensed into compact, rod-shaped chromosomes. Histone modifications play a key role in this process: They change significantly during condensation and regulate the conversion of chromatin.
For the first time, researchers have precisely tracked how molecular marks on DNA proteins change during cell division—and disproved a long-held assumption in the process.
An international research team led by Professor Axel Imhof at LMU’s Biomedical Center and Professor William Earnshaw (University of Edinburgh) has analyzed these changes during cell division with unprecedented precision. To this end, the researchers developed an innovative method that synchronizes the division of cell populations. They then employed high-resolution mass spectrometry to precisely record the changes in histone modifications during cell division. The findings are published in Molecular Cell.