Nociception refers to how our nerves respond to stimuli that trigger pain. Nerves in skin and other peripheral tissues – such as muscles and joints – that detect damaging stimuli are called nociceptors; they relay signals to the brain to initiate pain.
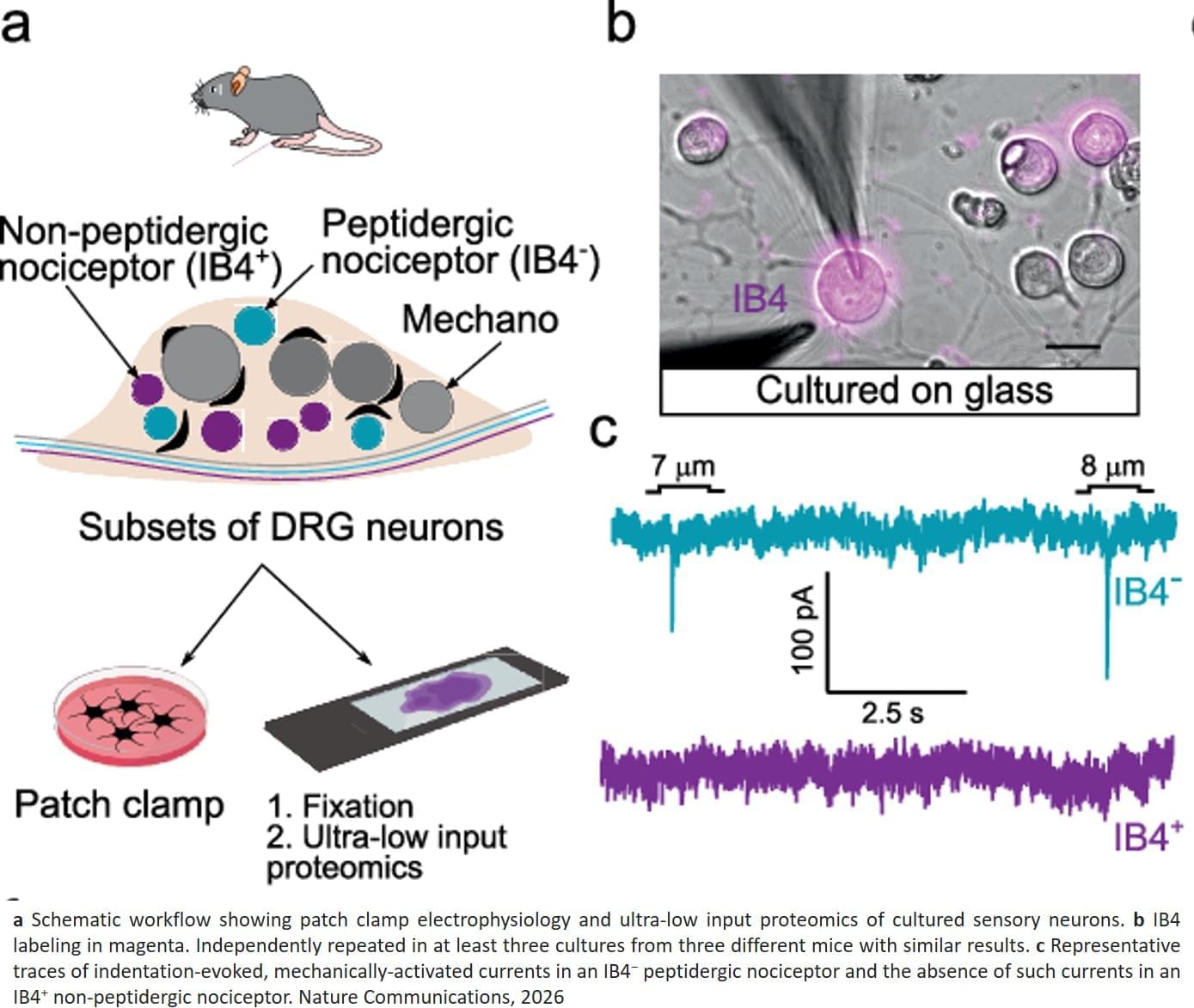
Using an electrophysiological method known as the patch-clamp technique, the team first identified and characterized two nociceptor subtypes – peptidergic and non-peptidergic – in the spinal ganglia of mice. Each of these subtypes respond differently to similar stimuli and may initiate pain of different quality and duration.
The researchers used around 50 neurons of each subtype to generate a specific protein map for each of the two cell types. Deep Visual Proteomics combines mass spectrometry with microscopy, artificial intelligence and robotics. The team have so far mainly used this methodology for proteome analyses of cancer cells. “We have now shown for the first time that it can also be applied to nerve cells,” another co-senior author says.
The team measured more than 6,000 proteins in these 50 neurons. A comparison with existing RNA data revealed that the transcriptome and proteome of the cells differ significantly in some cases – an indication that key functional processes only become visible at the protein level. “We provide a unique molecular map of pain-initiating neurons,” says the author. “It enables the identification of signaling pathways in these cells that have so far remained hidden.”
In an additional step, the authors wanted to understand which proteins sensitize nerve cells, contributing to chronic pain. They isolated both types of nociceptors from mouse dorsal root ganglia and exposed them to a molecule called Nerve Growth Factor (NGF), which is known to trigger chronic pain both animals and humans, such as in arthritis. Using Deep Visual Proteomics, the researchers were able to precisely identify the proteins produced after the cells were exposed to NGF.
“We identified several proteins that were present in higher levels in a subset of nociceptors following treatment with NGF. The higher levels of these proteins could be linked to long term pain associated with inflammation,” says the first author. One of the proteins, an enzyme called B3GNT2, stood out in particular. “When we knocked out the corresponding gene in the cells, the inflammation-induced hyperactivity of nociceptors was reduced. Fewer cells responded to mechanical stimulus,” the author says. In other words, the neurons had become less sensitive and would elicit much less pain. ScienceMission sciencenewshighlights.
One in five people worldwide suffers from chronic inflammatory pain. Meanwhile, about two thirds of those affected find little relief from existing pain medications; new therapeutic approaches are urgently needed. “We first must understand precisely how sensory nerve cells trigger pain at the molecular level — in other words, which proteins are involved,” says the senior author.
To unravel these molecular processes, two groups of researchers co-developed a method called Deep Visual Proteomics that makes it possible to determine the proteome — the complete set of proteins — of specific cells and to create maps detailing the spatial locations of individual proteins.
The researchers combined this technology with electrophysiological methods. This enabled them to first identify specific subtypes of pain neurons based on their function and then analyze their protein profiles. The result is a high-resolution molecular map of these nerve cells, which has been published in “Nature Communications.” The team also demonstrated how the technology can identify potential new drugs targets to treat chronic pain.