https://doi.org/10.1172/JCI199841 As part of the JCI’s Review Series on Neurodegeneration, Olivia Gautier, Thao P. Nguyen & Aaron D. Gitler explore the molecular basis for selective neuronal vulnerability and degeneration and summarize recent advances and applications of single-cell genomic approaches.
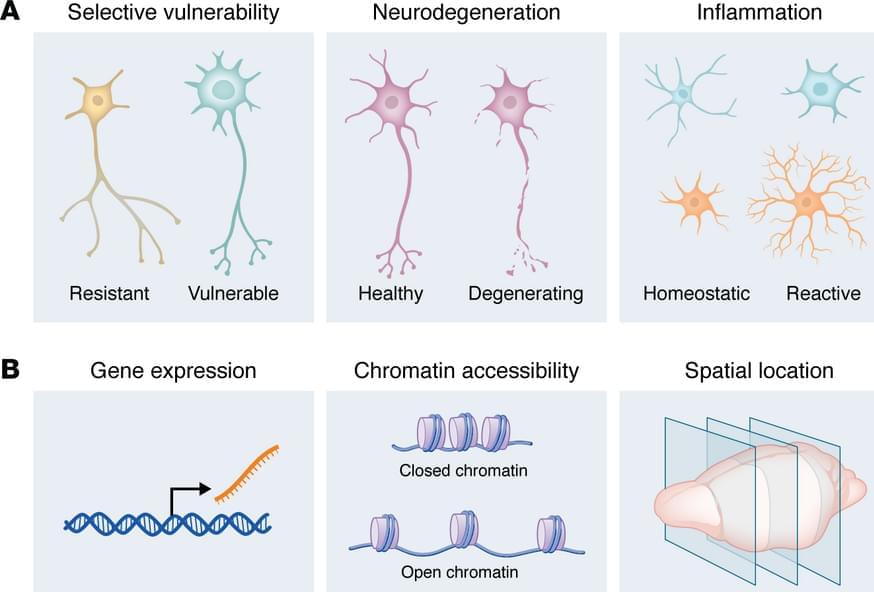
How do we decide whether we should choose single-cell or single-nucleus sequencing? This depends on sample types and biological applications. Single-cell sequencing is typically applied to fresh, readily dissociable tissues or cultured cells to study intact cell populations. Because it captures both cytoplasmic and nuclear transcripts, scRNA-seq provides a comprehensive view of cellular gene expression. However, tissue dissociation can induce stress-related transcriptional artifacts and introduce substantial cell-type bias. Large or fragile neurons are often lost during dissociation, whereas smaller cell types, such as astrocytes and oligodendrocytes, tend to be overrepresented. In contrast, single-nucleus sequencing is commonly used for frozen samples or for tissues that are difficult to dissociate, including the brain and spinal cord. Although fresh or fresh-frozen samples are typically used, snRNA-seq is compatible with formalin-fixed, paraffin-embedded (FFPE) samples, enabling the analysis of archived human specimens. A key limitation is that snRNA-seq does not capture cytoplasmic transcripts and is therefore biased toward nuclear, often premature, mRNA species.
Spatial transcriptomics does not require tissue dissociation and enables examination of cellular transcriptomes within their native tissue niches. Some spatial transcriptomic technologies are now compatible with FFPE samples, allowing analyses of preserved clinical specimens along with fixed-frozen and fresh-frozen samples. These technologies can be broadly classified into two main categories: imaging-based and sequencing-based (Figure 2B). Imaging-based approaches, like multiplexed error-robust fluorescence in situ hybridization (MERFISH), spatially resolved transcript amplicon readout mapping (STARmap), and 10x Genomics Xenium, rely on probe hybridization and multiplexed imaging to detect and visualize transcripts at high spatial resolution, often achieving single-cell or even subcellular resolution (17, 18). Although whole-transcriptome measurements are possible, MERFISH typically targets predefined gene panels due to the constraints of iterative hybridization and imaging. In contrast, sequencing-based approaches, including NanoString GeoMx and 10x Genomics Visium, capture RNA on spatially barcoded tissue slides or nanobeads followed by next-generation sequencing. These methods generally recover a broader range of transcripts than imaging-based approaches but, in most cases, do not yet achieve true single-cell resolution. Instead, they measure gene expression within spatial “spots” that encompass multiple cells and therefore rely on computational deconvolution to infer cell-type composition. Newer spatial transcriptomic methods, like spatial enhanced resolution omics sequencing (Stereo-seq) and reverse-padlock amplicon-encoding fluorescence in situ hybridization (RAEFISH), are approaching single-cell and single-molecule resolution (19 – 21).
In this Review, we summarize recent advances and applications of single-cell genomics approaches to study neurodegenerative disorders, including Alzheimer disease (AD), Parkinson disease (PD), amyotrophic lateral sclerosis (ALS), frontotemporal dementia (FTD), and Huntington disease (HD). We focus on how these approaches provide insight into the unique vulnerabilities of specific neuronal populations, define novel disease-associated cellular states, and reveal contributions of non-neuronal cells to disease pathogenesis. We then look to the future, envisioning how these technologies will empower genetic screens to uncover modifiers of neurodegeneration and new therapeutic targets.