Apr 26, 2024
More efficient molecular motor widens potential applications
Posted by Shailesh Prasad in categories: chemistry, nanotechnology
Light-driven molecular motors were first developed nearly 25 years ago at the University of Groningen, the Netherlands. This resulted in a shared Nobel Prize for Chemistry for Professor Ben Feringa in 2016. However, making these motors do actual work proved to be a challenge. A new paper from the Feringa lab, published in Nature Chemistry on 26 April, describes a combination of improvements that brings real-life applications closer.
First author Jinyu Sheng, now a postdoctoral researcher at the Institute of Science and Technology Austria (ISTA), adapted a “first generation” light-driven molecular motor during his Ph.D. studies in the Feringa laboratory. His main focus was to increase the efficiency of the motor molecule. “It is very fast, but only 2% of the photons that the molecule absorbs drive the rotary movement.”
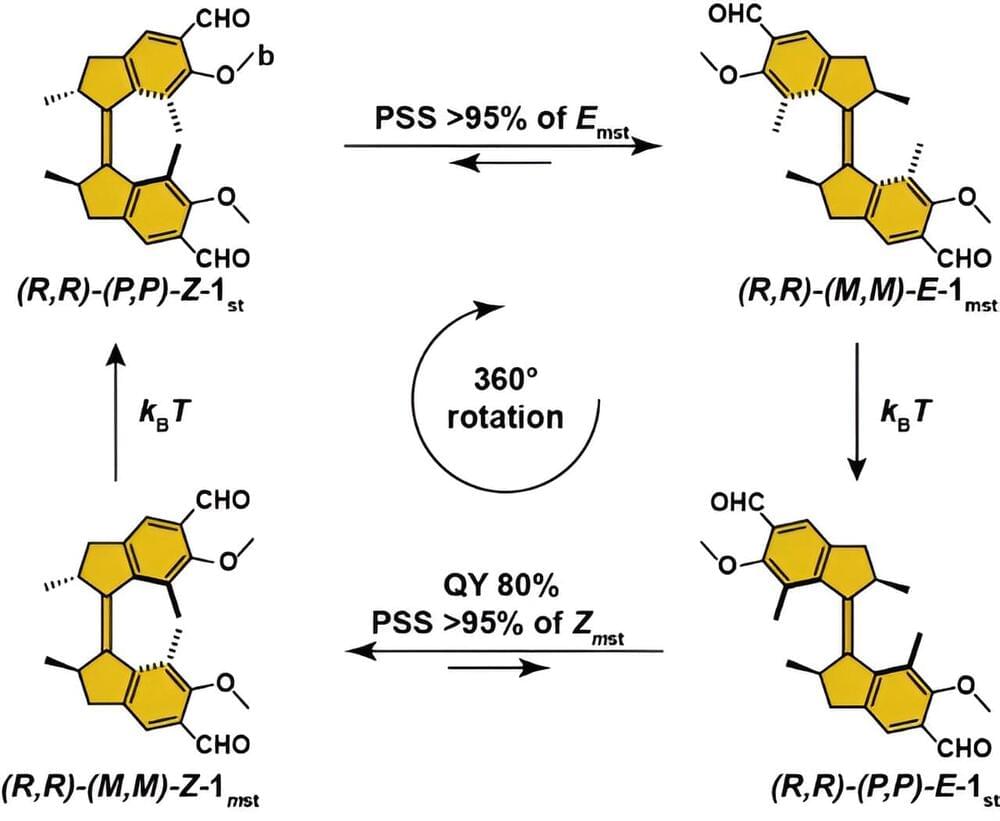
This poor efficiency can get in the way of real-life applications. “Besides, increased efficiency would give us better control of the motion,” adds Sheng. The rotary motion of Feringa’s molecular motor takes place in four steps: two of them are photochemical, while two are temperature-driven. The latter are unidirectional, but the photochemical steps cause an isomerization of the molecule that is usually reversible.