After three years of collecting scores of data on hundreds of stars, the ULLYSES (Ultraviolet Legacy Library of Young Stars as Essential Standards) survey conducted by NASA’s Hubble Space Telescope officially ended in December 2023, culminating in 220 total stars examined during the survey on data regarding their size, distance from Earth, temperature, chemical characteristics, and rotational speed. Additionally, ULYYSES also contains another 275 stars from the Hubble archive, providing researchers with several decades of new stellar data and holds the potential to help astronomers gain new insights into stellar formation and evolution throughout the universe.
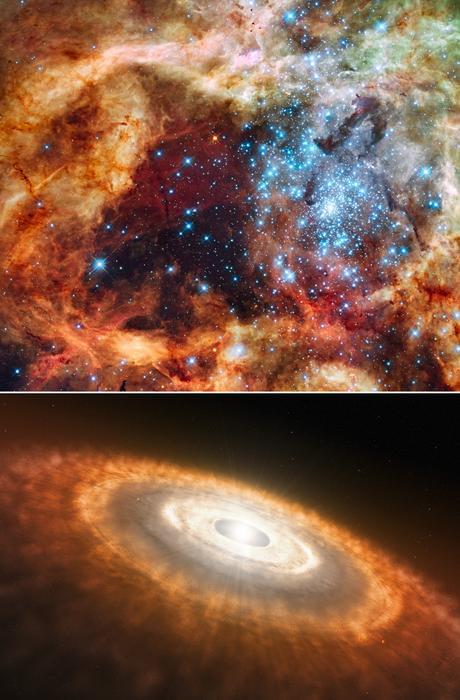
Hubble image of a star-forming region known as the Tarantula Nebula, which contains massive, young blue stars, which was observed during the ULYYSES survey (top panel). Artist’s illustration of a cooler, redder, young star smaller than our Sun that is still gathering material from its planet-forming disk (bottom panel). (Credit: NASA, ESA, STScI, Francesco Paresce (INAF-IASF Bologna), Robert O’Connell (UVA), SOC-WFC3, ESO)
“I believe the ULLYSES project will be transformative, impacting overall astrophysics – from exoplanets, to the effects of massive stars on galaxy evolution, to understanding the earliest stages of the evolving universe,” said Dr. Julia Roman-Duval, who is Implementation Team Lead for ULLYSES and an Associate Astronomer at the Space Telescope Science Institute (STScI). “Aside from the specific goals of the program, the stellar data can also be used in fields of astrophysics in ways we can’t yet imagine.”