AI language models, used to generate human-like text to power chatbots and create content, are also revolutionizing biology by treating complex biological data like a language. Language models are increasingly used, for example, to find patterns in DNA and proteins, to make predictions and speed research into biological complexity. A critical gap, however, is the lack of a method to estimate the reliability of these predictions.
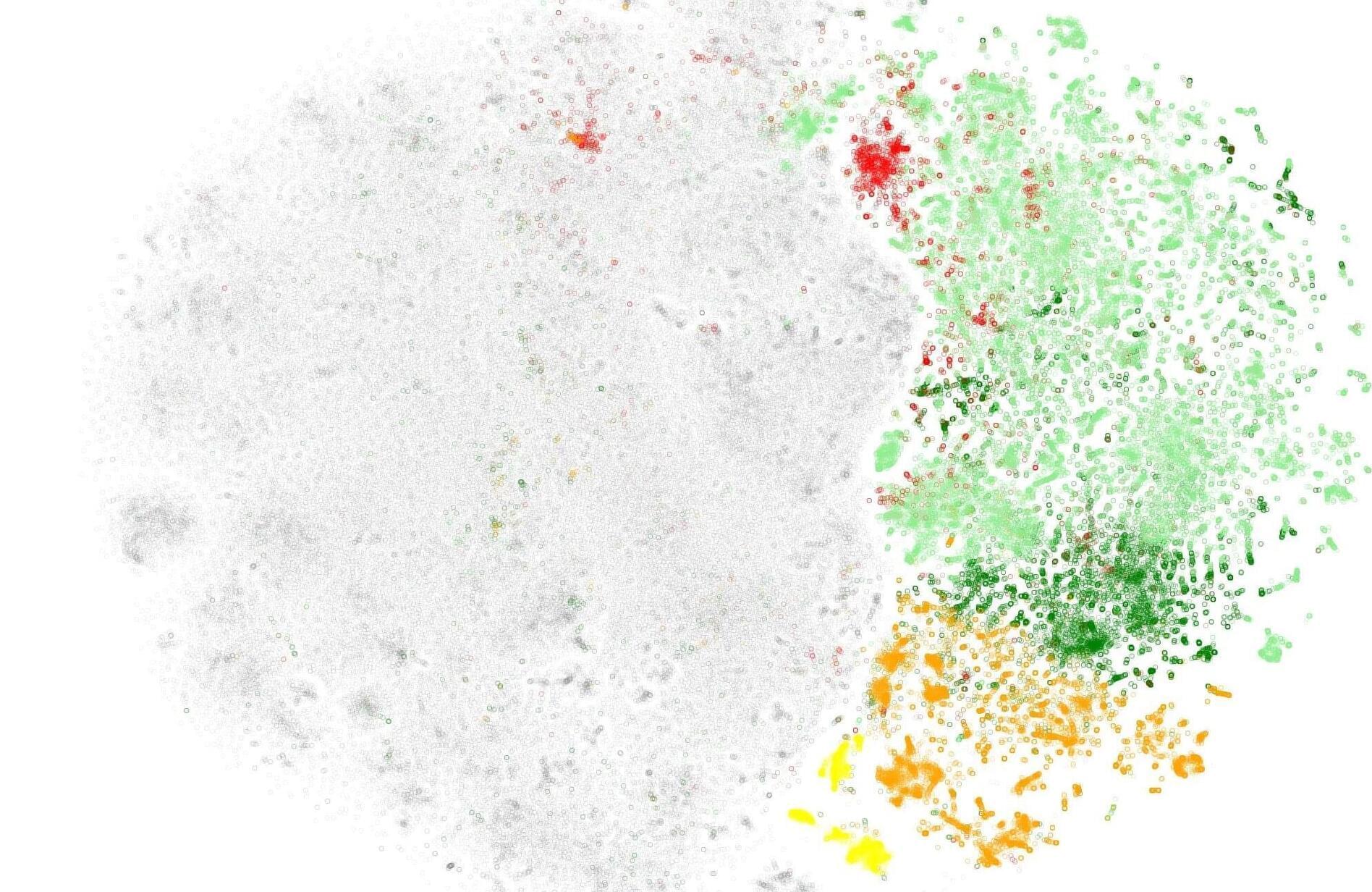
Computational biologists at Emory University have bridged this gap, developing a simple way to test the accuracy of a language model’s understanding of proteins. Nature Methods has published their system, which scores the reliability of a model’s predictions by comparing how it embeds (numerically codifies) synthetic random proteins versus proteins found in nature.
“To the best of our knowledge, our framework is the first generalized method to quantify protein sequence embedding reliability,” says Yana Bromberg, senior author of the paper and Emory professor of biology and computer science.